AFM observation of protein translocation mediated by one unit of SecYEG-SecA complex
- PMID: 39779699
- PMCID: PMC11711467
- DOI: 10.1038/s41467-024-54875-x
AFM observation of protein translocation mediated by one unit of SecYEG-SecA complex
Abstract
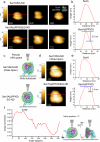
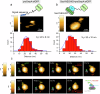
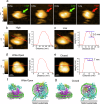
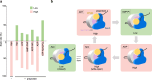
Protein translocation across cellular membranes is an essential and nano-scale dynamic process. In the bacterial cytoplasmic membrane, the core proteins in this process are a membrane protein complex, SecYEG, corresponding to the eukaryotic Sec61 complex, and a cytoplasmic protein, SecA ATPase. Despite more than three decades of extensive research on Sec proteins, from genetic experiments to cutting-edge single-molecule analyses, no study has visually demonstrated protein translocation. Here, we visualize the translocation, via one unit of a SecYEG-SecA-embedded nanodisc, of an unfolded substrate protein by high-speed atomic force microscopy (HS-AFM). Additionally, the uniform unidirectional distribution of nanodiscs on a mica substrate enables the HS-AFM image data analysis, revealing dynamic structural changes in the polypeptide-crosslinking domain of SecA between wide-open and closed states depending on nucleotides. The nanodisc-AFM approach will allow us to execute detailed analyses of Sec proteins as well as visualize nano-scale events of other membrane proteins.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures
References
-
- Rapoport, T. A., Li, L. & Park, E. Structural and mechanistic insights into protein translocation. Annu. Rev. Cell Dev. Biol.33, 369–390 (2017). - PubMed
-
- Tsirigotaki, A., De Geyter, J., Šoštaric, N., Economou, A. & Karamanou, S. Protein export through the bacterial Sec pathway. Nat. Rev. Microbiol.15, 21–36 (2017). - PubMed
-
- Tanaka, Y. et al. Crystal structures of SecYEG in lipidic cubic phase elucidate a precise resting and a peptide-bound state. Cell Rep.13, 1561–1568 (2015). - PubMed
-
- Economou, A. & Wickner, W. SecA promotes preprotein translocation by undergoing ATP-driven cycles of membrane insertion and deinsertion. Cell78, 835–843 (1994). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- JPMJKP23H2/MEXT | Japan Science and Technology Agency (JST)
- JPMJPR20E1/MEXT | Japan Science and Technology Agency (JST)
- hp230209, hp240215, hp240277/Ministry of Education, Culture, Sports, Science and Technology (MEXT)
- JPMXP1323015482/Ministry of Education, Culture, Sports, Science and Technology (MEXT)
- 24ZR1403800/Natural Science Foundation of Shanghai (Natural Science Foundation of Shanghai Municipality)
LinkOut - more resources
Full Text Sources
Miscellaneous

