DigiLoCS: A leap forward in predictive organ-on-chip simulations
- PMID: 39787162
- PMCID: PMC11717216
- DOI: 10.1371/journal.pone.0314083
DigiLoCS: A leap forward in predictive organ-on-chip simulations
Abstract
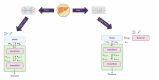
Digital twins, driven by data and mathematical modelling, have emerged as powerful tools for simulating complex biological systems. In this work, we focus on modelling the clearance on a liver-on-chip as a digital twin that closely mimics the clearance functionality of the human liver. Our approach involves the creation of a compartmental physiological model of the liver using ordinary differential equations (ODEs) to estimate pharmacokinetic (PK) parameters related to on-chip liver clearance. The objectives of this study were twofold: first, to predict human clearance values, and second, to propose a framework for bridging the gap between in vitro findings and their clinical relevance. The methodology integrated quantitative Organ-on-Chip (OoC) and cell-based assay analyses of drug depletion kinetics and is further enhanced by incorporating an OoC-digital twin model to simulate drug depletion kinetics in humans. The in vitro liver clearance for 32 drugs was predicted using a digital-twin model of the liver-on-chip and in vitro to in vivo extrapolation (IVIVE) was assessed using time series PK data. Three ODEs in the model define the drug concentrations in media, interstitium and intracellular compartments based on biological, hardware, and physicochemical information. A key issue in determining liver clearance appears to be the insufficient drug concentration within the intracellular compartment. The digital twin establishes a connection between the hardware chip structure and an advanced mapping of the underlying biology, specifically focusing on the intracellular compartment. Our modelling offers the following benefits: i) better prediction of intrinsic liver clearance of drugs compared to the conventional model and ii)explainability of behaviour based on physiological parameters. Finally, we illustrate the clinical significance of this approach by applying the findings to humans, utilising propranolol as a proof-of-concept example. This study stands out as the biggest cross-organ-on-chip platform investigation to date, systematically analysing and predicting human clearance values using data obtained from various in vitro liver-on-chip systems. Accurate prediction of in vivo clearance from in vitro data is important as inadequate understanding of the clearance of a compound can lead to unexpected and undesirable outcomes in clinical trials, ranging from underdosing to toxicity. Physiologically based pharmacokinetic (PBPK) model estimation of liver clearance is explored. The aim is to develop digital twins capable of determining better predictions of clinical outcomes, ultimately reducing the time, cost, and patient burden associated with drug development. Various hepatic in vitro systems are compared and their effectiveness for predicting human clearance is investigated. The developed tool, DigiLoCs, focuses explicitly on accurately describing complex biological processes within liver-chip systems. ODE-constrained optimisation is applied to estimate the clearance of compounds. DigiLoCs enable differentiation between active biological processes (metabolism) and passive processes (permeability and partitioning) by incorporating detailed information on compound-specific characteristics and hardware-specific data. These findings signify a significant stride towards more accurate and efficient drug development methodologies.
Copyright: © 2025 Aravindakshan et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures





References
-
- Denayer T, Stöhr T, Roy MV. Animal models in translational medicine: validation and prediction. European Journal of Molecular & Clinical Medicine. 2014;2(1):5. doi: 10.1016/j.nhtm.2014.08.001 - DOI
-
- Thelen K, Coboeken K, Willmann S, Burghaus R, Dressman JB, Lippert J. Evolution of a detailed physiological model to simulate the gastrointestinal transit and absorption process in humans, part 1: oral solutions. Journal of Pharmaceutical Sciences. 2011;100(12):5324–5345. doi: 10.1002/jps.22726 - DOI - PubMed
-
- Obach RS. Prediction of human clearance of twenty-nine drugs from hepatic microsomal intrinsic clearance data: an examination of in vitro half-life approach and nonspecific binding to microsomes. Drug Metabolism and Disposition. 1999;27(11):1350–1359. - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous

