Host-microbe multiomic profiling identifies distinct COVID-19 immune dysregulation in solid organ transplant recipients
- PMID: 39794319
- PMCID: PMC11723965
- DOI: 10.1038/s41467-025-55823-z
Host-microbe multiomic profiling identifies distinct COVID-19 immune dysregulation in solid organ transplant recipients
Abstract
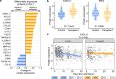
Coronavirus disease 2019 (COVID-19) poses significant risks for solid organ transplant recipients, who have atypical but poorly characterized immune responses to infection. We aim to understand the host immunologic and microbial features of COVID-19 in transplant recipients by leveraging a prospective multicenter cohort of 86 transplant recipients age- and sex-matched with 172 non-transplant controls. We find that transplant recipients have higher nasal SARS-CoV-2 viral abundance and impaired viral clearance, and lower anti-spike IgG levels. In addition, transplant recipients exhibit decreased plasmablasts and transitional B cells, and increased senescent T cells. Blood and nasal transcriptional profiling demonstrate unexpected upregulation of innate immune signaling pathways and increased levels of several proinflammatory serum chemokines. Severe disease in transplant recipients, however, is characterized by a less robust induction of pro-inflammatory genes and chemokines. Together, our study reveals distinct immune features and altered viral dynamics in solid organ transplant recipients.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: F.K. has the following financial interests: The Icahn School of Medicine at Mount Sinai has filed patent applications relating to SARS-CoV-2 serological assays, NDV-based SARS-CoV-2 vaccines, influenza virus vaccines, and influenza virus therapeutics which list Florian Krammer as co-inventor (Patent title and number: Influenza Virus Vaccines and Uses Thereof (Chimeric HA 2) 9,371,366; Influenza Virus Vaccines and Uses Thereof (Chimeric HA 1) 10,131,695; Influenza Virus Vaccines and Uses Thereof (Chimeric HA 2) 2934581; Influenza Virus Vaccines and Uses Thereof (Chimeric HA 2) 9,968,670; Influenza Virus Vaccines and Uses Thereof (Chimeric HA 2) 10,137,189, Influenza Virus Vaccines and Uses Thereof (Chimeric HA 2) 10,583,188; Influenza Virus Vaccines and Uses Thereof (Chimeric HA 1) EP2758075; Influenza Virus Vaccination Regimens (Neuraminidase) 10,736,956; Anti-Influenza B Virus Neuraminidase Antibodies and Uses Thereof 11254733; Influenza Virus Hemagluttinin Proteins and Uses Thereof (Mosaic) 7237344). Mount Sinai has spun out a company, Kantaro, to market serological tests for SARS-CoV-2 and another company, Castlevax, to develop SARS-CoV-2 vaccines. F.K. is a co-founder and scientific advisory board member of Castlevax. F.K. has consulted for Merck, Curevac, Seqirus, and Pfizer and is currently consulting for 3rd Rock Ventures, GSK, Gritstone, and Avimex. The Krammer laboratory is also collaborating with Dynavax on influenza vaccine development. R.R.M. has a Leadership Councilor role 2018-2021 for the Society of Leukocyte Biology. O.L. has received support as a speaker for presentation regarding the Coronavirus pandemic from Midsized Bank Coalition of Americ (MBCA) and Moody’s Analytics. N.G.R. has research grants from Pfizer, Merck, Sanofi, Quidel, Immorna, Vaccine Company, and Lilly, serves on safety committees for ICON and EMMES and the advisory boards of Moderna, Seqirus, Pfizer, and Sanofi, and is a paid safety consultant for ICON, CyanVac and EMMES. The remaining authors declare no competing interests.
Figures







Update of
-
Host-Microbe Multi-omic Profiling Identifies a Unique Program of COVID-19 Inflammatory Dysregulation in Solid Organ Transplant Recipients.Res Sq [Preprint]. 2023 Dec 20:rs.3.rs-3621844. doi: 10.21203/rs.3.rs-3621844/v1. Res Sq. 2023. Update in: Nat Commun. 2025 Jan 10;16(1):586. doi: 10.1038/s41467-025-55823-z. PMID: 38196658 Free PMC article. Updated. Preprint.
References
-
- World Health Organization. WHO Coronavirus (COVID-19) Dashboard. https://covid19.who.int (2023).
Publication types
MeSH terms
Substances
Grants and funding
- U19 AI090023/AI/NIAID NIH HHS/United States
- 5R01HL155418/U.S. Department of Health & Human Services | NIH | National Heart, Lung, and Blood Institute (NHLBI)
- U54 AI142766/AI/NIAID NIH HHS/United States
- U19 AI057229/AI/NIAID NIH HHS/United States
- U19 AI062629/AI/NIAID NIH HHS/United States
- U19 AI077439/AI/NIAID NIH HHS/United States
- U19 AI118610/AI/NIAID NIH HHS/United States
- R01 AI104870/AI/NIAID NIH HHS/United States
- U19 AI125357/AI/NIAID NIH HHS/United States
- R01 AI145835/AI/NIAID NIH HHS/United States
- I01 BX005023/BX/BLRD VA/United States
- U19 AI128913/AI/NIAID NIH HHS/United States
- U19 AI118608/AI/NIAID NIH HHS/United States
- UL1 TR001863/TR/NCATS NIH HHS/United States
- P01 AI153559/AI/NIAID NIH HHS/United States
- R01 AI135803/AI/NIAID NIH HHS/United States
- U19 AI089992/AI/NIAID NIH HHS/United States
- R01 HL155418/HL/NHLBI NIH HHS/United States
- U19 AI128910/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

