Erwinia phage Asesino is a nucleus-forming phage that lacks PhuZ
- PMID: 39799172
- PMCID: PMC11724907
- DOI: 10.1038/s41598-024-64095-4
Erwinia phage Asesino is a nucleus-forming phage that lacks PhuZ
Abstract
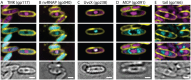
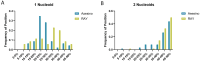
As nucleus-forming phages become better characterized, understanding their unifying similarities and unique differences will help us understand how they occupy varied niches and infect diverse hosts. All identified nucleus-forming phages fall within the Chimalliviridae family and share a core genome of 68 unique genes including chimallin, the major nuclear shell protein. A well-studied but non-essential protein encoded by many nucleus-forming phages is PhuZ, a tubulin homolog which aids in capsid migration, nucleus rotation, and nucleus positioning. One clade that represents 24% of all currently known chimalliviruses lacks a PhuZ homolog. Here we show that Erwinia phage Asesino, one member of this PhuZ-less clade, shares a common overall replication mechanism with other characterized nucleus-forming phages despite lacking PhuZ. We show that Asesino replicates via a phage nucleus that encloses phage DNA and partitions proteins in the nuclear compartment and cytoplasm in a manner similar to previously characterized nucleus-forming phages. Consistent with a lack of PhuZ, however, we did not observe active positioning or rotation of the phage nucleus within infected cells. These data show that some nucleus-forming phages have evolved to replicate efficiently without PhuZ, providing an example of a unique variation in the nucleus-based replication pathway.
© 2024. The Author(s).
Conflict of interest statement
Competing interests: J.P. has an equity interest in Linnaeus Bioscience Incorporated and receives income. The terms of this arrangement have been reviewed and approved by the University of California, San Diego, in accordance with its conflict-of-interest policies.
Figures



Update of
-
Asesino: a nucleus-forming phage that lacks PhuZ.bioRxiv [Preprint]. 2024 May 11:2024.05.10.593592. doi: 10.1101/2024.05.10.593592. bioRxiv. 2024. Update in: Sci Rep. 2025 Jan 11;15(1):1692. doi: 10.1038/s41598-024-64095-4. PMID: 38766163 Free PMC article. Updated. Preprint.
Similar articles
-
Asesino: a nucleus-forming phage that lacks PhuZ.bioRxiv [Preprint]. 2024 May 11:2024.05.10.593592. doi: 10.1101/2024.05.10.593592. bioRxiv. 2024. Update in: Sci Rep. 2025 Jan 11;15(1):1692. doi: 10.1038/s41598-024-64095-4. PMID: 38766163 Free PMC article. Updated. Preprint.
-
Identifying the core genome of the nucleus-forming bacteriophage family and characterization of Erwinia phage RAY.Cell Rep. 2023 May 30;42(5):112432. doi: 10.1016/j.celrep.2023.112432. Epub 2023 Apr 28. Cell Rep. 2023. PMID: 37120812 Free PMC article.
-
Identifying the core genome of the nucleus-forming bacteriophage family and characterization of Erwinia phage RAY.bioRxiv [Preprint]. 2023 Feb 25:2023.02.24.529968. doi: 10.1101/2023.02.24.529968. bioRxiv. 2023. Update in: Cell Rep. 2023 May 30;42(5):112432. doi: 10.1016/j.celrep.2023.112432. PMID: 36865095 Free PMC article. Updated. Preprint.
-
The intricate organizational strategy of nucleus-forming phages.Curr Opin Microbiol. 2024 Jun;79:102457. doi: 10.1016/j.mib.2024.102457. Epub 2024 Apr 6. Curr Opin Microbiol. 2024. PMID: 38581914 Free PMC article. Review.
-
The Phage Nucleus and PhuZ Spindle: Defining Features of the Subcellular Organization and Speciation of Nucleus-Forming Jumbo Phages.Front Microbiol. 2021 Jul 13;12:641317. doi: 10.3389/fmicb.2021.641317. eCollection 2021. Front Microbiol. 2021. PMID: 34326818 Free PMC article. Review.
References
-
- Malone, L. M. et al. A jumbo phage that forms a nucleus-like structure evades CRISPR-Cas DNA targeting but is vulnerable to type III RNA-based immunity. Nat. Microbiol.5, 48–55 (2020). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

