This is a preprint.
Immune targets for schistosomiasis control identified by a genome-wide association study of African snail vectors
- PMID: 39801518
- PMCID: PMC11722531
- DOI: 10.21203/rs.3.rs-5656395/v1
Immune targets for schistosomiasis control identified by a genome-wide association study of African snail vectors
Update in
-
Genes linked to schistosome resistance identified in a genome-wide association study of African snail vectors.Nat Commun. 2025 Jul 27;16(1):6918. doi: 10.1038/s41467-025-61760-8. Nat Commun. 2025. PMID: 40715059 Free PMC article.
Abstract
Schistosomiasis, a neglected tropical disease, is transmitted by freshwater snails. Interruption of transmission will require novel vector-focused interventions. We performed a genome-wide association study of African snails, Biomphalaria sudanica, exposed to Schistosoma mansoni in an endemic area of high transmission in Kenya. Two snail genomic regions, SudRes1 and SudRes2, were significantly associated with snail immunity to schistosomes. SudRes1 includes receptor-like protein tyrosine phosphatases while SudRes2 includes a class of leucine-rich repeat-containing G-protein coupled receptors, both comprising diverse extracellular binding domains suggestive of host-pathogen interaction. Resistant and susceptible haplotypes show numerous coding differences including presence/absence of entire genes. No loci previously tied to schistosome resistance in neotropical snail species showed any association with compatibility suggesting that loci involved in the resistance of African vectors are distinct. Snail ancestry was also strongly correlated with parasite compatibility. These results will inform future efforts to predict and manipulate immunity of a major schistosome vector.
Conflict of interest statement
Competing Interest Statement: The authors declare no competing interests
Figures



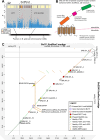
References
-
- WHO, Global report on neglected tropical diseases 2024 (2024).
-
- Hay S. I., et al. , Global, regional, and national disability-adjusted life-years (DALYs) for 333 diseases and injuries and healthy life expectancy (HALE) for 195 countries and territories, 1990–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet 390, 1260–1344 (2017). - PMC - PubMed
-
- WHO, “WHO guideline on control and elimination of human schistosomiasis” (2022). - PubMed
-
- WHO, “Ending the neglect to attain the Sustainable Development Goals – A road map for neglected tropical diseases 2021–2030” (2020).
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

