This is a preprint.
Combined statistical-biophysical modeling links ion channel genes to physiology of cortical neuron types
- PMID: 39803528
- PMCID: PMC11722265
- DOI: 10.1101/2023.03.02.530774
Combined statistical-biophysical modeling links ion channel genes to physiology of cortical neuron types
Update in
-
Combined statistical-biophysical modeling links ion channel genes to physiology of cortical neuron types.Patterns (N Y). 2025 Aug 5;6(10):101323. doi: 10.1016/j.patter.2025.101323. eCollection 2025 Oct 10. Patterns (N Y). 2025. PMID: 41142913 Free PMC article.
Abstract
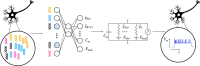
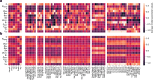
Neural cell types have classically been characterized by their anatomy and electrophysiology. More recently, single-cell transcriptomics has enabled an increasingly fine genetically defined taxonomy of cortical cell types, but the link between the gene expression of individual cell types and their physiological and anatomical properties remains poorly understood. Here, we develop a hybrid modeling approach to bridge this gap. Our approach combines statistical and mechanistic models to predict cells' electrophysiological activity from their gene expression pattern. To this end, we fit biophysical Hodgkin-Huxley-based models for a wide variety of cortical cell types using simulation-based inference, while overcoming the challenge posed by the mismatch between the mathematical model and the data. Using multimodal Patch-seq data, we link the estimated model parameters to gene expression using an interpretable sparse linear regression model. Our approach recovers specific ion channel gene expressions as predictive of biophysical model parameters including ion channel densities, directly implicating their mechanistic role in determining neural firing.
Conflict of interest statement
Declaration of Interests The authors declare no competing interests.
Figures






References
-
- Zeng H. & Sanes J. R. Neuronal cell-type classification: challenges, opportunities and the path forward. Nat. Rev. Neurosci. 18, 530–546 (2017). - PubMed
-
- Douglas R. J., Martin K. A. et al. Neuronal circuits of the neocortex. Annu. review neuroscience 27, 419–451 (2004). - PubMed
-
- Markram H. et al. Reconstruction and simulation of neocortical microcircuitry. Cell 163, 456–492 (2015). - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
