Human Hair Follicle Mesenchymal Stem Cell-Derived Exosomes Attenuate UVB-Induced Photoaging via the miR-125b-5p/TGF-β1/Smad Axis
- PMID: 39807308
- PMCID: PMC11725759
- DOI: 10.34133/bmr.0121
Human Hair Follicle Mesenchymal Stem Cell-Derived Exosomes Attenuate UVB-Induced Photoaging via the miR-125b-5p/TGF-β1/Smad Axis
Abstract
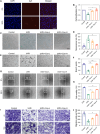
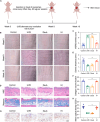
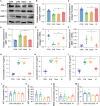
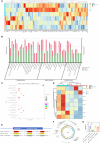
Cutaneous photoaging, induced by chronic exposure to ultraviolet (UV) radiation, typically manifests as alterations in both the physical appearance and functional properties of the skin and may predispose individuals to cancer development. Recent studies have demonstrated the reparative potential of exosomes derived from mesenchymal stem cells in addressing skin damage, while specific reports highlight their efficacy in ameliorating skin photoaging. However, the precise role of exosomes derived from human hair follicle mesenchymal stem cells (HFMSC-Exos) in the context of cutaneous photoaging remains largely unexplored. We successfully isolated HFMSC-Exos using the ultracentrifugation technique. In cellular experiments, we assessed the migration of human dermal fibroblasts (HDFs) through scratch and transwell assays, evaluated the angiogenesis of human umbilical vein endothelial cells through angiogenesis assays, and examined the expression levels of collagen and matrix metalloproteinase 1 (MMP-1) using Western blotting and quantitative reverse transcription polymerase chain reaction. Furthermore, we established a nude mouse model of photoaging to observe wrinkle formation on the dorsal surface of the animals, as well as to assess dermal thickness and collagen fiber generation through histological staining. Ultimately, we performed RNA sequencing on skin tissues from mice before and after treatment to elucidate the relevant underlying mechanisms. Our findings revealed that HFMSC-Exos effectively enhanced the migration and proliferation of HDFs and upregulated the expressions of transforming growth factor-β1 (TGF-β1), p-Smad2/p-Smad3, collagen type 1, and collagen type 3 while concurrently down-regulating MMP-1 levels in HDFs. Additionally, mice in the HFMSC-Exo group showed quicker wrinkle healing and increased collagen production. HFMSC-Exos miR-125b-5p was demonstrated to reduce skin photoaging by increasing profibrotic levels via TGF-β1 expression. UV-irradiated HDFs and photoaged nude mouse skin showed low TGF-β1 expressions, whereas overexpression of TGF-β1 in HDFs increased collagen type 1, collagen type 3, and p-Smad2/p-Smad3 expressions while decreasing MMP-1 expression. HDFs overexpressing TGF-β1 produced more collagen and altered the Smad pathway. This study demonstrated, both in vitro and in vivo, that HFMSC-Exos increased collagen formation, promoted HDF cell proliferation and migration, and reversed the senescence of UV-irradiated HDFs. TGF-β1 was identified as a target of HFMSC-Exos miR-125b-5p, which controls photoaging via regulating the Smad pathway. The antiphotoaging capabilities of HFMSC-Exos may occur via the miR-125b-5p/TGF-β1/Smad axis, suggesting a promising therapeutic approach for treating skin photoaging.
Copyright © 2025 Hong Cui et al.
Conflict of interest statement
Competing interests: The authors declare that they have no competing interests.
Figures


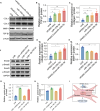
References
-
- Stratigos AJ, Katsambas AD. The role of topical retinoids in the treatment of photoaging. Drugs. 2005;65(8):1061–1072. - PubMed
-
- Ichihashi M, Ando H. The maximal cumulative solar UVB dose allowed to maintain healthy and young skin and prevent premature photoaging. Exp Dermatol. 2014;23(Suppl 1):43–46. - PubMed
-
- Battie C, Jitsukawa S, Bernerd F, Del Bino S, Marionnet C, Verschoore M. New insights in photoaging, UVA induced damage and skin types. Exp Dermatol. 2014;23(Suppl 1):7–12. - PubMed
-
- Botchkarev VA, Kishimoto J. Molecular control of epithelial–mesenchymal interactions during hair follicle cycling. J Investig Dermatol Symp Proc. 2003;8(1):46–55. - PubMed
LinkOut - more resources
Full Text Sources

