LIN28B-mediated PI3K/AKT pathway activation promotes metastasis in colorectal cancer models
- PMID: 39808497
- PMCID: PMC11996871
- DOI: 10.1172/JCI186035
LIN28B-mediated PI3K/AKT pathway activation promotes metastasis in colorectal cancer models
Abstract
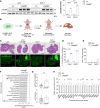
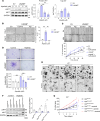
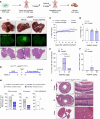
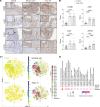
Colorectal cancer (CRC) remains a leading cause of cancer death because of metastatic spread. LIN28B is overexpressed in 30% of CRCs and promotes metastasis, yet its mechanisms remain unclear. In this study, we genetically modified CRC cell lines to overexpress LIN28B, resulting in enhanced PI3K/AKT pathway activation and liver metastasis in mice. We developed genetically modified mouse models with constitutively active Pik3ca that form intestinal tumors progressing to liver metastases with an intact immune system, addressing the limitations of previous Pik3ca-mutant models, including long tumor latency, mixed histology, and lack of distant metastases. The PI3Kα-specific inhibitor alpelisib reduced migration and invasion in vitro and metastasis in vivo. We present a comprehensive analysis of vertical inhibition of the PI3K/AKT pathway in CRC using the FDA-approved drugs alpelisib and capivasertib (an AKT inhibitor) in combination with LY2584702 (a ribosomal protein S6 kinase inhibitor) in CRC cell lines and mouse- and patient-derived organoids. Tissue microarrays from patients with CRC verified that LIN28B and PI3K/AKT pathway activation correlate with CRC progression. These findings highlight the critical role of the LIN28B-mediated PI3K/AKT pathway in CRC metastasis, the therapeutic potential of targeted inhibition, and the promise of patient-derived organoids in precision medicine in metastatic CRC.
Keywords: Colorectal cancer; Gastroenterology; Mouse models; Oncogenes; Oncology.
Conflict of interest statement
Figures




References
-
- American Cancer Society. Survival Rates for Colorectal Cancer. https://www.cancer.org/cancer/types/colon-rectal-cancer/detection-diagno... Updated January 29, 2024. Accessed September 1, 2024.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

