The members of zinc finger-homeodomain (ZF-HD) transcription factors are associated with abiotic stresses in soybean: insights from genomics and expression analysis
- PMID: 39810081
- PMCID: PMC11730174
- DOI: 10.1186/s12870-024-06028-x
The members of zinc finger-homeodomain (ZF-HD) transcription factors are associated with abiotic stresses in soybean: insights from genomics and expression analysis
Abstract
Background: Zinc finger homeodomain (ZF-HD) belongs to the plant-specific transcription factor (TF) family and is widely involved in plant growth, development and stress responses. Despite their importance, a comprehensive identification and analysis of ZF-HD genes in the soybean (Glycine max) genome and their possible roles under abiotic stress remain unexplored.
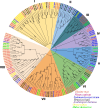
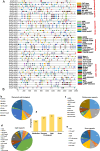
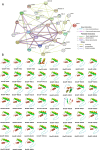
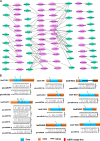
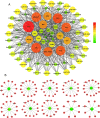
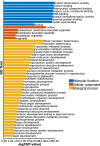
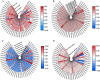
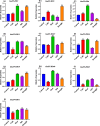
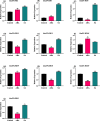
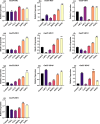
Results: In this study, 51 ZF-HD genes were identified in the soybean genome that were unevenly distributed on 17 chromosomes. All GmZF-HD genes contained a conserved ZF-HD_dimer domain and had diverse physicochemical features. Furthermore, the GmZF-HD gene structures exhibited 3 to 10 conserved motifs, and most of them showed intronless gene structures. Phylogenetic analysis categorized them into eight major groups with the highest closeness to dicots including Brassica rapa and Malus domestica. The cis-element analysis recognized plant growth and development (10%), phytohormones (31%) and stress-responsive (59%) elements. Synteny analysis identified 73 segmental and 1 tandem duplicated genes that underwent purifying selection. The collinearity analysis revealed that GmZF-HD genes showed higher homology with dicot species, indicating common ancestors with close evolutionary relationships. A total of 94 gma-miRNAs from 41 diverse miRNA families were identified, targeting 40 GmZF-HD genes, with GmZF-HD6 being most targeted by 7 miRNAs, and gma-miR4993 emerging as the dominant miRNA family. Different TFs including ERF, LBD, BBR-BPC and MYB, etc., were predicted in all 51 GmZF-HD genes upstream regions and visualized in the network. Expression profiling through RNA-Seq showed diverse expressions of GmZF-HD genes in different tissues including seeds, roots, shoots and leaves under diverse conditions. Further, the qRT-PCR analysis demonstrated that all tested GmZF-HD genes were significantly induced in soybean leaves, mainly the GmZF-HD5/6/13/39 and GmZF-HD45 genes were significantly upregulated (2.5 to 8.8 folds) under the tested stress treatments compared to control, highlighting their potential roles in response to stresses in soybean.
Conclusion: Overall, this study reveals comprehensive insights into the ZF-HD genes in soybeans and provides a valuable contribution towards functional studies for soybean improvement under stress conditions.
Keywords: Abiotic stress; Gene expressions; Micro-RNA; Soybean; Synteny; ZF-HD.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: All experimental research on plants, including the collection of plant materials has been used by following the relevant institutional, national, and international guidelines and legislation. Competing interests: The authors declare no competing interests.
Figures


Similar articles
-
Pathogen-induced binding of the soybean zinc finger homeodomain proteins GmZF-HD1 and GmZF-HD2 to two repeats of ATTA homeodomain binding site in the calmodulin isoform 4 (GmCaM4) promoter.Nucleic Acids Res. 2007;35(11):3612-23. doi: 10.1093/nar/gkm273. Epub 2007 May 7. Nucleic Acids Res. 2007. PMID: 17485478 Free PMC article.
-
ZF-HD gene family in rapeseed (Brassica napus L.): genome-wide identification, phylogeny, evolutionary expansion and expression analyses.BMC Genomics. 2024 Dec 5;25(1):1181. doi: 10.1186/s12864-024-11102-7. BMC Genomics. 2024. PMID: 39639240 Free PMC article.
-
Identification and Transcriptional Analysis of Zinc Finger-Homeodomain (ZF-HD) Family Genes in Cucumber.Biochem Genet. 2021 Aug;59(4):884-901. doi: 10.1007/s10528-021-10036-z. Epub 2021 Feb 8. Biochem Genet. 2021. PMID: 33554320
-
Zinc Finger-Homeodomain and Mini Zinc Finger proteins are key players in plant growth and responses to environmental stresses.J Exp Bot. 2022 Aug 11;73(14):4662-4673. doi: 10.1093/jxb/erac194. J Exp Bot. 2022. PMID: 35536651 Review.
-
Diverse roles of MYB transcription factors in plants.J Integr Plant Biol. 2025 Mar;67(3):539-562. doi: 10.1111/jipb.13869. Epub 2025 Feb 27. J Integr Plant Biol. 2025. PMID: 40013511 Review.
References
-
- Gavili E, Moosavi AA, Haghighi AAK. Does biochar mitigate the adverse effects of drought on the agronomic traits and yield components of soybean? Ind Crops Prod. 2019;128:445–54.
-
- Raza A, Bashir S, Khare T, Karikari B, Copeland RG, Jamla M, Abbas S, Charagh S, Nayak SN, Djalovic I. Temperature-smart plants: A new horizon with omics-driven plant breeding. Physiol Plant. 2024;176(1):e14188.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

