Genomic variation responding to artificial selection on different lines of Pekin duck
- PMID: 39813863
- PMCID: PMC11783388
- DOI: 10.1016/j.psj.2025.104785
Genomic variation responding to artificial selection on different lines of Pekin duck
Abstract
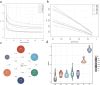
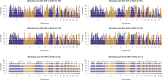
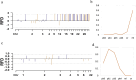
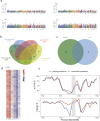
Understanding the genomic variation in Pekin duck under artificial selection is important for improving the utilization of duck genetic resources. Here, the genomic changes in Pekin duck were analyzed by using the genome resequencing data from 96 individual samples, including 2 conservation populations and 4 breeding populations with different breeding backgrounds. The population structure, runs of homozygosity (ROH), effective population number (Ne), and other genetic parameters were analyzed. The breeding populations showed lower genetic diversity compared to the conservation populations. Maple Leaf duck and Cherry Valley duck retained low genetic diversity compared to other breeding populations, with Cherry Valley duck showing the lowest diversity and the highest inbreeding coefficient. This suggested that Cherry Valley and Maple Leaf ducks have undergone intensive selection compared to other breeding populations. By the analysis of runs of homozygosity (ROHs), some genes (e.g., IGF1R) associated with growth traits were identified. By the analysis of the selection signal, strong selection characteristics in certain genomic regions during the breeding of Peking duck across different selected lines were observed. In addition, copy number variations (CNVs) in Pekin duck populations were analyzed. Six regions of interest were identified, containing RPA1, DOT1L, SLC25A42, RALYL, TRPA1, and IGFBP2. Furthermore, the allele frequency distribution of these genes showed significant differences between breeding populations and conservation populations, indicating that these candidate genes could have undergone strong selection pressure during long-term selection for improved production. These findings contribute to a deeper understanding of the distinct evolutionary processes in Pekin ducks under artificial selection and provide valuable insights for future breeding strategies.
Keywords: Genetic diversity; Intensive selection; Pekin duck; Weight.
Copyright © 2025. Published by Elsevier Inc.
Conflict of interest statement
Declaration of competing interest None.
Figures


Similar articles
-
Resequencing of a Pekin duck breeding population provides insights into the genomic response to short-term artificial selection.Gigascience. 2023 Mar 20;12:giad016. doi: 10.1093/gigascience/giad016. Epub 2023 Mar 27. Gigascience. 2023. PMID: 36971291 Free PMC article.
-
Genome-wide association analysis reveals 6 copy number variations associated with the number of cervical vertebrae in Pekin ducks.Front Cell Dev Biol. 2022 Nov 10;10:1041088. doi: 10.3389/fcell.2022.1041088. eCollection 2022. Front Cell Dev Biol. 2022. PMID: 36438573 Free PMC article.
-
Whole-genome selection signature differences between Chaohu and Ji'an red ducks.BMC Genomics. 2024 May 27;25(1):522. doi: 10.1186/s12864-024-10339-6. BMC Genomics. 2024. PMID: 38802792 Free PMC article.
-
A comparative analysis reveals the genomic diversity among 8 Muscovy duck populations.G3 (Bethesda). 2024 Jul 8;14(7):jkae112. doi: 10.1093/g3journal/jkae112. G3 (Bethesda). 2024. PMID: 38789099 Free PMC article.
-
Leveraging Whole-Genome Resequencing to Uncover Genetic Diversity and Promote Conservation Strategies for Ruminants in Asia.Animals (Basel). 2025 Mar 13;15(6):831. doi: 10.3390/ani15060831. Animals (Basel). 2025. PMID: 40150358 Free PMC article. Review.
References
-
- Alberto F.J., Boyer F., Orozco-terWengel P., Streeter I., Servin B., de Villemereuil P., Benjelloun B., Librado P., Biscarini F., Colli L., Barbato M., Zamani W., Alberti A., Engelen S., Stella A., Joost S., Ajmone-Marsan P., Negrini R., Orlando L., Rezaei H.R., Naderi S., Clarke L., Flicek P., Wincker P., Coissac E., Kijas J., Tosser-Klopp G., Chikhi A., Bruford M.W., Taberlet P., Pompanon F. Convergent genomic signatures of domestication in sheep and goats. Nat. Commun. 2018;9:813. - PMC - PubMed
-
- Aldosary M., Baselm S., Abdulrahim M., Almass R., Alsagob M., AlMasseri Z., Huma R., AlQuait L., Al-Shidi T., Al-Obeid E., AlBakheet A., Alahideb B., Alahaidib L., Qari A., Taylor R.W., Colak D., AlSayed M.D., Kaya N. SLC25A42-associated mitochondrial encephalomyopathy: Report of additional founder cases and functional characterization of a novel deletion. JIMD Rep. 2021;60:75–87. - PMC - PubMed
-
- Ashton M. Crowood; 2014. Domestic Duck.
-
- Axelsson E., Ratnakumar A., Arendt M.L., Maqbool K., Webster M.T., Perloski M., Liberg O., Arnemo J.M., Hedhammar A., Lindblad-Toh K. The genomic signature of dog domestication reveals adaptation to a starch-rich diet. Nature. 2013;495:360–364. - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous

