Novel Quinazoline Derivatives Inhibit Splicing of Fungal Group II Introns
- PMID: 39824511
- PMCID: PMC11851433
- DOI: 10.1021/acschembio.4c00631
Novel Quinazoline Derivatives Inhibit Splicing of Fungal Group II Introns
Abstract
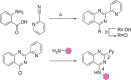
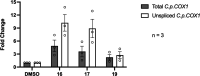
We report the discovery of small molecules that target the RNA tertiary structure of self-splicing group II introns and display potent antifungal activity against yeasts, including the major public health threat Candida parapsilosis. High-throughput screening efforts against a yeast group II intron resulted in an inhibitor class which was then synthetically optimized for enhanced inhibitory activity and antifungal efficacy. The most highly refined compounds in this series display strong, gene-specific antifungal activity against C. parapsilosis. This work demonstrates the utility of combining advanced RNA screening methodologies with medicinal chemistry pipelines to identify high-affinity ligands targeting RNA tertiary structures with important roles in human health and disease.
Conflict of interest statement
The authors declare the following competing financial interest(s): A.M.P. is a founder of IntronX, an RNA-targeted antifungal drug discovery company. The quinazoline compounds described in this work are covered by filings from Yale University.
Figures


Similar articles
-
Small molecules that target group II introns are potent antifungal agents.Nat Chem Biol. 2018 Dec;14(12):1073-1078. doi: 10.1038/s41589-018-0142-0. Epub 2018 Oct 15. Nat Chem Biol. 2018. PMID: 30323219 Free PMC article.
-
Group I Intron as a Potential Target for Antifungal Compounds: Development of a Trans-Splicing High-Throughput Screening Strategy.Molecules. 2023 May 31;28(11):4460. doi: 10.3390/molecules28114460. Molecules. 2023. PMID: 37298936 Free PMC article.
-
Identification of an Exceptionally Long Intron in the HAC1 Gene of Candida parapsilosis.mSphere. 2018 Nov 7;3(6):e00532-18. doi: 10.1128/mSphere.00532-18. mSphere. 2018. PMID: 30404939 Free PMC article.
-
Candida parapsilosis: from Genes to the Bedside.Clin Microbiol Rev. 2019 Feb 27;32(2):e00111-18. doi: 10.1128/CMR.00111-18. Print 2019 Mar 20. Clin Microbiol Rev. 2019. PMID: 30814115 Free PMC article. Review.
-
Candida and candidaemia. Susceptibility and epidemiology.Dan Med J. 2013 Nov;60(11):B4698. Dan Med J. 2013. PMID: 24192246 Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

