Dual RNA sequencing of a co-culture model of Pseudomonas aeruginosa and human 2D upper airway organoids
- PMID: 39824906
- PMCID: PMC11742674
- DOI: 10.1038/s41598-024-82500-w
Dual RNA sequencing of a co-culture model of Pseudomonas aeruginosa and human 2D upper airway organoids
Abstract
Pseudomonas aeruginosa is a Gram-negative bacterium that is notorious for airway infections in cystic fibrosis (CF) subjects. Bacterial quorum sensing (QS) coordinates virulence factor expression and biofilm formation at population level. Better understanding of QS in the bacterium-host interaction is required. Here, we set up a new P. aeruginosa infection model, using 2D upper airway nasal organoids that were derived from 3D organoids. Using dual RNA-sequencing, we dissected the interaction between organoid epithelial cells and WT or QS-mutant P. aeruginosa strains. Since only a single healthy individual and a single CF subject were used as donors for the organoids, conclusions about CF-specific effects could not be deduced. However, P. aeruginosa induced epithelial inflammation, whereas QS signaling did not affect the epithelial airway cells. Conversely, the epithelium influenced infection-related processes of P. aeruginosa, including QS-mediated regulation. Comparison of our model with samples from the airways of CF subjects indicated that our model recapitulates important aspects of infection in vivo. Hence, the 2D airway organoid infection model is relevant and may help to reduce the future burden of P. aeruginosa infections in CF.
Keywords: Pseudomonas aeruginosa; 2D co-culture; Airway organoids; Dual RNA-sequencing; Infection model; Quorum sensing.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: HC is the head of Pharma Research and Early Development at Roche, Basel and holds several patents related to organoids technology. HC’s full disclosure is given at https://www.uu.nl/staff/JCClevers/. The other authors declare no conflict of interest.
Figures
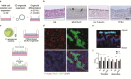





References
-
- Forbes, E. & Abu-Sbaih, R. Cystic fibrosis. Nat. Rev. Dis. Primers. 10.1038/nrdp.2015.49 (2015). - DOI
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

