This is a preprint.
Transcriptional Integration of Meiotic Prophase I Progression and Early Oocyte Differentiation
- PMID: 39829852
- PMCID: PMC11741336
- DOI: 10.1101/2025.01.06.631470
Transcriptional Integration of Meiotic Prophase I Progression and Early Oocyte Differentiation
Abstract
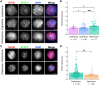
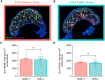
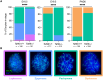
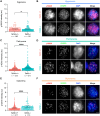
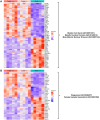
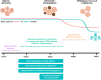
Female reproductive senescence results from the regulated depletion of a finite pool of oocytes called the ovarian reserve. This pool of oocytes is initially established during fetal development, but the oocytes that comprise it must remain quiescent for decades until they are activated during maturation in adulthood. In order for developmentally competent oocytes to populate the ovarian reserve they must successfully initiate both meiosis and oogenesis. As the factors that regulate the timing and fidelity of these early events remain elusive, we assessed the precise function and timing of the transcriptional regulator TAF4b during meiotic prophase I progression in mouse fetal oocytes. Compared to matched controls, E14.5 Taf4b-deficient oocytes enter meiosis I in a timely manner however, their subsequent progression through the pachytene-to-diplotene transition of meiotic prophase I is compromised. Moreover, this disruption of meiotic progression is associated with the reduced ability of Taf4b-deficient oocytes to repair double-strand DNA breaks. Transcriptional profiling of Taf4b-deficient oocytes reveals that between E16.5 and E18.5 these oocytes fail to coordinate the reduction of meiotic gene expression and the induction of oocyte differentiation genes. These studies reveal that TAF4b promotes the formation of the ovarian reserve in part by orchestrating the timely transition to meiosis I arrest and oocyte differentiation, which are often perceived as separate events.
Figures


Similar articles
-
TAF4b transcription networks regulating early oocyte differentiation.Development. 2022 Feb 1;149(3):dev200074. doi: 10.1242/dev.200074. Epub 2022 Feb 9. Development. 2022. PMID: 35043944 Free PMC article.
-
Molecular analysis of the effects of steroid hormones on mouse meiotic prophase I progression.Reprod Biol Endocrinol. 2019 Dec 2;17(1):105. doi: 10.1186/s12958-019-0548-x. Reprod Biol Endocrinol. 2019. PMID: 31791345 Free PMC article.
-
Arrest at the diplotene stage of meiotic prophase I is delayed by progesterone but is not required for primordial follicle formation in mice.Reprod Biol Endocrinol. 2016 Dec 5;14(1):82. doi: 10.1186/s12958-016-0218-1. Reprod Biol Endocrinol. 2016. PMID: 27919266 Free PMC article.
-
Regulation of Meiotic Prophase One in Mammalian Oocytes.Front Cell Dev Biol. 2021 May 20;9:667306. doi: 10.3389/fcell.2021.667306. eCollection 2021. Front Cell Dev Biol. 2021. PMID: 34095134 Free PMC article. Review.
-
The molecular regulatory mechanisms of meiotic arrest and resumption in Oocyte development and maturation.Reprod Biol Endocrinol. 2023 Oct 2;21(1):90. doi: 10.1186/s12958-023-01143-0. Reprod Biol Endocrinol. 2023. PMID: 37784186 Free PMC article. Review.
References
-
- Abby E., Tourpin S., Ribeiro J., Daniel K., Messiaen S., Moison D., Guerquin J., Gaillard J. C., Armengaud J., Langa F., Toth A., Martini E., & Livera G. (2016). Implementation of meiosis prophase I programme requires a conserved retinoid-independent stabilizer of meiotic transcripts. Nat Commun, 7, 10324. 10.1038/ncomms10324 - DOI - PMC - PubMed
-
- Alexander A. K., Rice E. J., Lujic J., Simon L. E., Tanis S., Barshad G., Zhu L., Lama J., Cohen P. E., & Danko C. G. (2023). A-MYB and BRDT-dependent RNA Polymerase II pause release orchestrates transcriptional regulation in mammalian meiosis. Nat Commun, 14(1), 1753. 10.1038/s41467-023-37408-w - DOI - PMC - PubMed
-
- Anderson E. L., Baltus A. E., Roepers-Gajadien H. L., Hassold T. J., de Rooij D. G., van Pelt A. M., Page D. C., Baudat F., Imai Y., & de Massy B. (2008). Stra8 and its inducer, retinoic acid, regulate meiotic initiation in both spermatogenesis and oogenesis in mice Meiotic recombination in mammals: localization and regulation. Proc Natl Acad Sci U S A, 105(39), 14976–14980. 10.1073/pnas.080729710510.1038/nrg3573 - DOI - PMC - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
