This is a preprint.
Serum, Cell-Free, HPV-Human DNA Junction Detection and HPV Typing for Predicting and Monitoring Cervical Cancer Recurrence
- PMID: 39830266
- PMCID: PMC11741462
- DOI: 10.1101/2024.09.16.24313343
Serum, Cell-Free, HPV-Human DNA Junction Detection and HPV Typing for Predicting and Monitoring Cervical Cancer Recurrence
Abstract
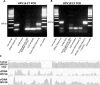
Almost all cervical cancers are caused by human papillomaviruses (HPVs). In most cases, HPV DNA is integrated into the human genome. We found that tumor-specific, HPV-human DNA junctions are detectable in serum cell-free DNA of a fraction of cervical cancer patients at the time of initial treatment and/or at six months following treatment. Retrospective analysis revealed these junctions were more frequently detectable in women in whom the cancer later recurred. We also found that cervical cancers caused by HPV types outside of phylogenetic clade α9 had a higher recurrence frequency than those caused by α9 types in both our study and The Cancer Genome Atlas cervical cancer database, despite the higher prevalence of α9 types including HPV16 in cervical cancer. Thus, HPV-human DNA junction detection in serum cell-free DNA and HPV type determination in tumor tissue may help predict recurrence risk. Screening serum cell-free DNA for junctions may also offer an unambiguous, non-invasive means to monitor absence of recurrence following treatment.
Keywords: DNA integration; HPV; cancer recurrence; cell-free DNA; cervical cancer; cfDNA; human papilloma virus.
Figures




Similar articles
-
[Association between high risk human papillomavirus DNA load and cervical lesions in different infection status].Zhonghua Zhong Liu Za Zhi. 2018 Jun 23;40(6):475-480. doi: 10.3760/cma.j.issn.0253-3766.2018.06.015. Zhonghua Zhong Liu Za Zhi. 2018. PMID: 29936777 Chinese.
-
Multiplex Identification of Human Papillomavirus 16 DNA Integration Sites in Cervical Carcinomas.PLoS One. 2013 Jun 18;8(6):e66693. doi: 10.1371/journal.pone.0066693. Print 2013. PLoS One. 2013. PMID: 23824673 Free PMC article.
-
Liquid biopsy of HPV DNA in cervical cancer.J Clin Virol. 2019 May;114:32-36. doi: 10.1016/j.jcv.2019.03.005. Epub 2019 Mar 12. J Clin Virol. 2019. PMID: 30913520
-
Biology and pathological associations of the human papillomaviruses: a review.Malays J Pathol. 1998 Jun;20(1):1-10. Malays J Pathol. 1998. PMID: 10879257 Review.
-
Cellular and molecular alterations in human epithelial cells transformed by recombinant human papillomavirus DNA.Crit Rev Oncog. 1993;4(4):337-60. Crit Rev Oncog. 1993. PMID: 8394744 Review.
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
