Epigenetic regulation of HOXA2 expression affects tumor progression and predicts breast cancer patient survival
- PMID: 39833374
- PMCID: PMC11982354
- DOI: 10.1038/s41418-024-01430-2
Epigenetic regulation of HOXA2 expression affects tumor progression and predicts breast cancer patient survival
Abstract
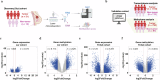
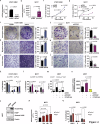
Accumulating evidence suggests that genetic and epigenetic biomarkers hold potential for enhancing the early detection and monitoring of breast cancer (BC). Epigenetic alterations of the Homeobox A2 (HOXA2) gene have recently garnered significant attention in the clinical management of various malignancies. However, the precise role of HOXA2 in breast tumorigenesis has remained elusive. To address this point, we conducted high-throughput RNA sequencing and DNA methylation array studies on laser-microdissected human BC samples, paired with normal tissue samples. Additionally, we performed comprehensive in silico analyses using large public datasets: TCGA and METABRIC. The diagnostic performance of HOXA2 was calculated by means of receiver operator characteristic curves. Its prognostic significance was assessed through immunohistochemical studies and Kaplan-Meier Plotter database interrogation. Moreover, we explored the function of HOXA2 and its role in breast carcinogenesis through in silico, in vitro, and in vivo investigations. Our work revealed significant hypermethylation and downregulation of HOXA2 in human BC tissues. Low HOXA2 expression correlated with increased BC aggressiveness and unfavorable patient survival outcomes. Suppression of HOXA2 expression significantly heightened cell proliferation, migration, and invasion in BC cells, and promoted tumor growth in mice. Conversely, transgenic HOXA2 overexpression suppressed these cellular processes and promoted apoptosis of cancer cells. Interestingly, a strategy of pharmacological demethylation successfully restored HOXA2 expression in malignant cells, reducing their neoplastic characteristics. Bioinformatics analyses, corroborated by in vitro experimentations, unveiled a novel implication of HOXA2 in the lipid metabolism of BC. Specifically, depletion of HOXA2 leaded to a concomitantly decreased expression of PPARγ and its target CIDEC, a master regulator of lipid droplet (LD) accumulation, thereby resulting in reduced LD abundance in BC cells. In summary, our study identifies HOXA2 as a novel prognosis-relevant tumor suppressor in the mammary gland.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: FDEDP, JGP, VDM, GK, FS, and MCM are listed as co-inventors on an international patent (“Methods for diagnosing, prognosing and managing treatment of breast cancer”; WO 2021/170777 A1). GK has been holding research contracts with Daiichi Sankyo, Eleor, Kaleido, Lytix Pharma, PharmaMar, Osasuna Therapeutics, Samsara Therapeutics, Sanofi, Tollys, and Vascage. GK has been consulting for Reithera. GK is on the Board of Directors of the Bristol Myers Squibb Foundation France. GK is a scientific co-founder of everImmune, Osasuna Therapeutics, Samsara Therapeutics, and Therafast Bio. GK is in the scientific advisory boards of Hevolution, Institut Servier, and Longevity Vision Funds. GK is the inventor of patents covering therapeutic targeting of aging, cancer, cystic fibrosis, and metabolic disorders. GK’s wife, Laurence Zitvogel, has held research contracts with Glaxo Smyth Kline, Incyte, Lytix, Kaleido, Innovate Pharma, Daiichi Sankyo, Pilege, Merus, Transgene, 9 m, Tusk, and Roche, was on the on the Board of Directors of Transgene, is a co-founder of everImmune, and holds patents covering the treatment of cancer and the therapeutic manipulation of the microbiota. GK’s brother, Romano Kroemer, was an employee of Sanofi and now consults for Boehringer-Ingelheim. The funders had no role in the design of the study; in the writing of the manuscript, or in the decision to publish the results. All the other authors declare no conflict of interest. Ethics approval and consent to participate: Human tissue samples have been used in agreement with regulatory and bioethics rules of the Istituto Nazionale Tumori - Fondazione G. Pascale Ethics Committee (protocol number 3 of 03/25/2009, Naples, Italy) [14]. All patients provided their written informed consent for the research purpose of these clinical materials according to the tenets of the Helsinki Declaration. Animal experiments were performed in compliance with the EU Directive 63/2010 and the local Ethical Committee (“C. Darwin” registered at the French Ministry of Research, protocol #51068).
Figures




References
-
- Harbeck N, Penault-Llorca F, Cortes J, Gnant M, Houssami N, Poortmans P, et al. Breast cancer. Nat Rev Dis Prim. 2019;5:1–31. - PubMed
-
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clinicians. 2020;70:7–30. - PubMed
-
- Loibl S, Poortmans P, Morrow M, Denkert C, Curigliano G. Breast cancer. Lancet. 2021;397:1750–69. - PubMed
-
- Waks AG, Winer EP. Breast cancer treatment: a review. JAMA. 2019;321:288–300. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials

