GOPhage: protein function annotation for bacteriophages by integrating the genomic context
- PMID: 39838963
- PMCID: PMC11751364
- DOI: 10.1093/bib/bbaf014
GOPhage: protein function annotation for bacteriophages by integrating the genomic context
Abstract
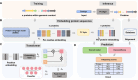
Bacteriophages are viruses that target bacteria, playing a crucial role in microbial ecology. Phage proteins are important in understanding phage biology, such as virus infection, replication, and evolution. Although a large number of new phages have been identified via metagenomic sequencing, many of them have limited protein function annotation. Accurate function annotation of phage proteins presents several challenges, including their inherent diversity and the scarcity of annotated ones. Existing tools have yet to fully leverage the unique properties of phages in annotating protein functions. In this work, we propose a new protein function annotation tool for phages by leveraging the modular genomic structure of phage genomes. By employing embeddings from the latest protein foundation models and Transformer to capture contextual information between proteins in phage genomes, GOPhage surpasses state-of-the-art methods in annotating diverged proteins and proteins with uncommon functions by 6.78% and 13.05% improvement, respectively. GOPhage can annotate proteins lacking homology search results, which is critical for characterizing the rapidly accumulating phage genomes. We demonstrate the utility of GOPhage by identifying 688 potential holins in phages, which exhibit high structural conservation with known holins. The results show the potential of GOPhage to extend our understanding of newly discovered phages.
Keywords: bacteriophages; genomic contextual information; protein function annotation; protein large language model.
© The Author(s) 2025. Published by Oxford University Press.
Figures



Similar articles
-
Improving viral annotation with artificial intelligence.mBio. 2024 Oct 16;15(10):e0320623. doi: 10.1128/mbio.03206-23. Epub 2024 Sep 4. mBio. 2024. PMID: 39230289 Free PMC article. Review.
-
Phage Genome Annotation Using the RAST Pipeline.Methods Mol Biol. 2018;1681:231-238. doi: 10.1007/978-1-4939-7343-9_17. Methods Mol Biol. 2018. PMID: 29134599
-
Essential Steps in Characterizing Bacteriophages: Biology, Taxonomy, and Genome Analysis.Methods Mol Biol. 2018;1681:197-215. doi: 10.1007/978-1-4939-7343-9_15. Methods Mol Biol. 2018. PMID: 29134597
-
Comparative Analyses of Bacteriophage Genomes.Methods Mol Biol. 2024;2802:427-453. doi: 10.1007/978-1-0716-3838-5_14. Methods Mol Biol. 2024. PMID: 38819567
-
Ecology, Structure, and Evolution of Shigella Phages.Annu Rev Virol. 2020 Sep 29;7(1):121-141. doi: 10.1146/annurev-virology-010320-052547. Epub 2020 May 11. Annu Rev Virol. 2020. PMID: 32392456 Free PMC article. Review.
Cited by
-
Characterization of holins, the membrane proteins of coliphage ASEC2201: a genomewide in silico approach.Front Microbiol. 2025 Jul 9;16:1550594. doi: 10.3389/fmicb.2025.1550594. eCollection 2025. Front Microbiol. 2025. PMID: 40703241 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

