The Epstein-Barr virus deubiquitinase BPLF1 regulates stress-induced ribosome UFMylation and reticulophagy
- PMID: 39842454
- PMCID: PMC12013442
- DOI: 10.1080/15548627.2024.2440846
The Epstein-Barr virus deubiquitinase BPLF1 regulates stress-induced ribosome UFMylation and reticulophagy
Abstract
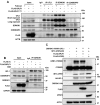
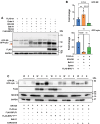
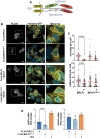
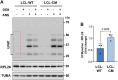
The synthesis of membrane and secreted proteins is safeguarded by an endoplasmic reticulum-associated ribosome quality control (ER-RQC) that promotes the disposal of defective translation products by the proteasome or via a lysosome-dependent pathway involving the degradation of portions of the ER by macroautophagy (reticulophagy). The UFMylation of RPL26 on ER-stalled ribosomes is essential for activating the ER-RQC and reticulophagy. Here, we report that the viral deubiquitinase (vDUB) encoded in the N-terminal domain of the Epstein-Barr virus (EBV) large tegument protein BPLF1 hinders the UFMylation of RPL26 on ribosomes that stall at the ER, promotes the stabilization of ER-RQC substrates, and inhibits reticulophagy. The vDUB did not act as a de-UFMylase or interfere with the UFMylation of the ER membrane protein CYB5R3 by the UFL1 ligase. Instead, it copurified with ribosomes in sucrose gradients and abrogated a ZNF598- and LTN1-independent ubiquitination event required for RPL26 UFMylation. Physiological levels of BPLF1 impaired the UFMylation of RPL26 in productively EBV-infected cells, pointing to an important role of the enzyme in regulating the translation quality control that allows the efficient synthesis of viral proteins and the production of infectious virus.Abbreviation: BPLF1, BamH1 P fragment left open readingframe-1; CDK5RAP3, CDK5regulatory subunit associated protein 3; ChFP, mCherry fluorescent protein; DDRGK1, DDRGKdomain containing 1; EBV, Epstein-Barr virus; eGFP, enhancedGFP; ER-RQC, endoplasmicreticulum-associated ribosome quality control; LCL, EBV-carryinglymphoblastoid cell line; GFP, green fluorescent protein; RQC, ribosome quality control; SRP, signal recognition particle; UFM1, ubiquitin fold modifier 1; UFL1, UFM1 specific ligase 1.
Keywords: EBV; Macroautophagy; Ribosome; UFM1; viral DUB.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures





References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
