Detection of Insertion/Deletions (InDel) Within Five Clock Genes and Their Associations with Growth Traits in Four Chinese Sheep Breeds
- PMID: 39852913
- PMCID: PMC11769414
- DOI: 10.3390/vetsci12010039
Detection of Insertion/Deletions (InDel) Within Five Clock Genes and Their Associations with Growth Traits in Four Chinese Sheep Breeds
Abstract
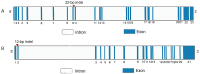
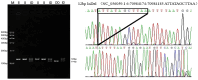
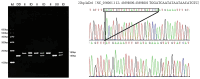
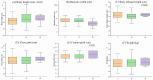
Organisms have the capacity to detect day-night fluctuations through oscillators regulated by circadian clock genes, which are crucial for regulating various biological processes. Numerous studies have demonstrated a marked association between these genes and various growth traits of sheep. This study identified polymorphisms at 23 potential loci within five clock genes in four Chinese sheep breeds. Only two polymorphic insertion/deletions (InDels) were detected in CLOCK and PER3 genes, respectively. The distribution of these two loci in four Chinese sheep breeds and their association with growth traits were further explored. A 12 bp deletion was found in the intron of the CLOCK gene (rs604230640), which was significantly associated with body height (p < 0.05), body oblique length (p < 0.05) and cannon girth (p < 0.05) in Hu sheep (HS). A 22 bp insertion in the intron of the PER3 gene (rs600537720) with a dominant genotype of insertion/insertion (II) was found to have a significant association with chest depth (p < 0.05) in Small-Tail Han sheep (STHS), tail width (p < 0.05) in Tong Sheep (TS), and in Lanzhou fat-tailed sheep (LFTS). In conclusion, this study has elucidated the polymorphisms of CLOCK and PER3 genes and has examined the influence of these two genes on the growth traits of sheep. Concurrently, the two molecular markers identified in CLOCK and PER3 could potentially serve in the marker-assisted selection of growing-related traits in local Chinese sheep breeds.
Keywords: association; circadian clock gene; growth traits; insertion/deletion (InDel); sheep.
Conflict of interest statement
The authors declare that no ethical conflicts of interest or significant financial support could have appeared edition fluence the work reported in this paper.
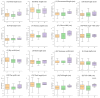
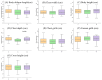
Figures
Similar articles
-
Identification and Analysis of InDel Variants in Key Hippo Pathway Genes and Their Association with Growth Traits in Four Chinese Sheep Breeds.Vet Sci. 2025 Mar 18;12(3):283. doi: 10.3390/vetsci12030283. Vet Sci. 2025. PMID: 40267010 Free PMC article.
-
Novel InDels of GHR, GHRH, GHRHR and Their Association with Growth Traits in Seven Chinese Sheep Breeds.Animals (Basel). 2020 Oct 15;10(10):1883. doi: 10.3390/ani10101883. Animals (Basel). 2020. PMID: 33076416 Free PMC article.
-
Two New Insertion/Deletion Variants of the PITX2 Gene and their Effects on Growth Traits in Sheep.Anim Biotechnol. 2018;29(4):276-282. doi: 10.1080/10495398.2017.1379415. Epub 2017 Dec 4. Anim Biotechnol. 2018. PMID: 29200321
-
Detection of key gene InDels in JAK/STAT pathway and their associations with growth traits in four Chinese sheep breeds.Gene. 2023 Dec 20;888:147750. doi: 10.1016/j.gene.2023.147750. Epub 2023 Aug 30. Gene. 2023. PMID: 37657690
-
Genetic variations in the sheep SIRT7 gene and their correlation with body size traits.Arch Anim Breed. 2019 Apr 16;62(1):189-197. doi: 10.5194/aab-62-189-2019. eCollection 2019. Arch Anim Breed. 2019. PMID: 31807629 Free PMC article.
References
-
- Hyten D.L., Cannon S.B., Song Q., Weeks N., Fickus E.W., Shoemaker R.C., Specht J.E., Farmer A.D., May G.D., Cregan P.B. High-Throughput SNP Discovery through Deep Resequencing of a Reduced Representation Library to Anchor and Orient Scaffolds in the Soybean Whole Genome Sequence. BMC Genom. 2010;11:38. doi: 10.1186/1471-2164-11-38. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

