Shigella sonnei and Shigella flexneri infection in Caenorhabditis elegans led to species-specific regulatory responses in the host and pathogen
- PMID: 39853209
- PMCID: PMC11893279
- DOI: 10.1099/mgen.0.001339
Shigella sonnei and Shigella flexneri infection in Caenorhabditis elegans led to species-specific regulatory responses in the host and pathogen
Abstract
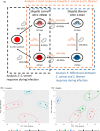
In recent decades, Shigella sonnei has surpassed Shigella flexneri as the leading cause of shigellosis, possibly due to species-specific differences in their transcriptomic responses. This study used dual RNA sequencing to analyse the transcriptomic responses of Caenorhabditis elegans and the two Shigella species at early (10 minutes) and late (24 hours) stages of infection. While the nematode defence response was downregulated during both Shigella infections, only infection by S. sonnei led to downregulation of sphingolipid metabolism, cadmium ion response and xenobiotic response in C. elegans. Furthermore, S. sonnei upregulates biofilm formation and energy generation/conservation during infection, acid resistance-related genes and biofilm regulators compared to S. flexneri. These findings highlight species-specific responses during C. elegans infection.
Keywords: Caenorhabditis elegans; Shigella flexneri; Shigella sonnei; dual-RNA sequencing; infection.
Conflict of interest statement
The authors declare that there are no conflicts of interest.
Figures



Similar articles
-
Shigella flexneri infection in Caenorhabditis elegans: cytopathological examination and identification of host responses.PLoS One. 2014 Sep 4;9(9):e106085. doi: 10.1371/journal.pone.0106085. eCollection 2014. PLoS One. 2014. PMID: 25187942 Free PMC article.
-
Transposon mutagenesis identifies acid resistance and biofilm genes as Shigella sonnei virulence factors in Caenorhabditis elegans infection.Biochem Biophys Res Commun. 2025 Mar 25;754:151546. doi: 10.1016/j.bbrc.2025.151546. Epub 2025 Mar 1. Biochem Biophys Res Commun. 2025. PMID: 40023989
-
Transcriptional response of Caenorhabditis elegans when exposed to Shigella flexneri.Genomics. 2020 Jan;112(1):774-781. doi: 10.1016/j.ygeno.2019.05.016. Epub 2019 May 21. Genomics. 2020. PMID: 31125598
-
Shigella sonnei: virulence and antibiotic resistance.Arch Microbiol. 2021 Jan;203(1):45-58. doi: 10.1007/s00203-020-02034-3. Epub 2020 Sep 14. Arch Microbiol. 2021. PMID: 32929595 Free PMC article. Review.
-
Shigella sonnei: epidemiology, evolution, pathogenesis, resistance and host interactions.Nat Rev Microbiol. 2025 May;23(5):303-317. doi: 10.1038/s41579-024-01126-x. Epub 2024 Nov 27. Nat Rev Microbiol. 2025. PMID: 39604656 Review.
References
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

