Development of a Polygenic Risk Score for Metabolic Dysfunction-Associated Steatotic Liver Disease Prediction in UK Biobank
- PMID: 39858580
- PMCID: PMC11765347
- DOI: 10.3390/genes16010033
Development of a Polygenic Risk Score for Metabolic Dysfunction-Associated Steatotic Liver Disease Prediction in UK Biobank
Abstract
Background: Metabolic dysfunction-associated steatotic liver disease (MASLD) is the leading cause of liver-related morbidity and mortality. Although the invasive liver biopsy remains the golden standard for MASLD diagnosis, Magnetic Resonance Imaging-derived Proton Density Fat Fraction (MRI-PDFF) is an accurate, non-invasive method for the assessment of treatment response. This study aimed at developing a Polygenic Risk Score (PRS) to improve MRI-PDFF prediction using UK Biobank data to assess an individual's genetic liability to MASLD.
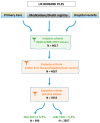
Methods: We iteratively sequestered 10% of MRI-PDFF samples as a validation set and split the rest of each dataset into base and target partitions, containing GWAS summary statistics and raw genotype data, respectively. PRSice2 was deployed to derive PRS candidates. Based on the frequency of SNP appearances along the PRS candidates, we generated different SNP sets according to variable frequency cutoffs. By applying the PRSs to the validation set, we identified the optimal SNP set, which was then applied to a Greek nonalcoholic fatty liver disease (NAFLD) study.
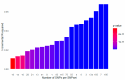
Results: Data from 3553 UK Biobank participants yielded 49 different SNP sets. After calculating the PRS on the validation set for every SNP set, an optimal PRS with 75 SNPs was selected (incremental R2 = 0.025, p-value = 0.00145). Interestingly, 43 SNPs were successfully mapped to MASLD-related known genes. The selected PRS could predict traits, like LDL cholesterol and diastolic blood pressure in the UK Biobank, as also disease outcome in the Greek NAFLD study.
Conclusions: Our findings provide strong evidence that PRS is a powerful prediction model for MASLD, while it can also be applied on populations of different ethnicity.
Keywords: UK Biobank; metabolic dysfunction-associated steatotic liver disease; polygenic risk score.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures




References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

