Methyl-dependent auto-regulation of the DNA N6-adenine methyltransferase AMT1 in the unicellular eukaryote Tetrahymena thermophila
- PMID: 39868535
- PMCID: PMC11760949
- DOI: 10.1093/nar/gkaf022
Methyl-dependent auto-regulation of the DNA N6-adenine methyltransferase AMT1 in the unicellular eukaryote Tetrahymena thermophila
Abstract
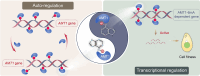
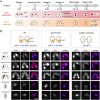
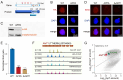
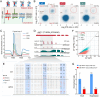
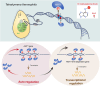
DNA N6-methyladenine (6mA) is a potential epigenetic mark involved in gene transcription in eukaryotes, yet the regulatory mechanism governing its methyltransferase (MTase) activity remains obscure. Here, we exploited the 6mA MTase AMT1 to elucidate its auto-regulation in the unicellular eukaryote Tetrahymena thermophila. The detailed endogenous localization of AMT1 in vegetative and sexual stages revealed a correlation between the 6mA reestablishment in the new MAC and the occurrence of zygotically expressed AMT1. Catalytically inactive AMT1 reduced 6mA level on the AMT1 gene and its expression, suggesting that AMT1 modulated its own transcription via 6mA. Furthermore, AMT1-dependent 6mA regulated the transcription of its target genes, thereby affecting cell fitness. Our findings unveil a positive feedback loop of transcriptional activation on the AMT1 gene and highlight the crucial role of AMT1-dependent 6mA in gene transcription.
© The Author(s) 2025. Published by Oxford University Press on behalf of Nucleic Acids Research.
Conflict of interest statement
None declared.
Figures


Similar articles
-
Identification and characterization of the de novo methyltransferases for eukaryotic N6-methyladenine (6mA).Sci Adv. 2025 May 16;11(20):eadq4623. doi: 10.1126/sciadv.adq4623. Epub 2025 May 14. Sci Adv. 2025. PMID: 40367178 Free PMC article.
-
Case Study of the Response of N6-Methyladenine DNA Modification to Environmental Stressors in the Unicellular Eukaryote Tetrahymena thermophila.mSphere. 2021 Jun 30;6(3):e0120820. doi: 10.1128/mSphere.01208-20. Epub 2021 May 28. mSphere. 2021. PMID: 34047647 Free PMC article.
-
Semiconservative transmission of DNA N 6-adenine methylation in a unicellular eukaryote.Genome Res. 2024 Jun 25;34(5):740-756. doi: 10.1101/gr.277843.123. Genome Res. 2024. PMID: 38744529 Free PMC article.
-
N6-methyladenine DNA modification in the unicellular eukaryotic organism Tetrahymena thermophila.Eur J Protistol. 2017 Apr;58:94-102. doi: 10.1016/j.ejop.2016.12.003. Epub 2016 Dec 23. Eur J Protistol. 2017. PMID: 28135687 Review.
-
The epigenetic roles of DNA N6-Methyladenine (6mA) modification in eukaryotes.Cancer Lett. 2020 Dec 1;494:40-46. doi: 10.1016/j.canlet.2020.08.025. Epub 2020 Aug 23. Cancer Lett. 2020. PMID: 32846189 Review.
Cited by
-
Identification and characterization of the de novo methyltransferases for eukaryotic N6-methyladenine (6mA).Sci Adv. 2025 May 16;11(20):eadq4623. doi: 10.1126/sciadv.adq4623. Epub 2025 May 14. Sci Adv. 2025. PMID: 40367178 Free PMC article.
-
Dual modes of DNA N6-methyladenine maintenance by distinct methyltransferase complexes.Proc Natl Acad Sci U S A. 2025 Jan 21;122(3):e2413037121. doi: 10.1073/pnas.2413037121. Epub 2025 Jan 15. Proc Natl Acad Sci U S A. 2025. PMID: 39813249 Free PMC article.
References
MeSH terms
Substances
Grants and funding
- 32200399/National Natural Science Foundation of China
- ZR2024ZD40/Natural Science Foundation of Shandong Province
- SDAST2024QTA008/Young Talent of Lifting engineering for Science and Technology in Shandong, China
- 202441014/Fundamental Research Funds for the Central Universities
- LSKJ202203203/Laoshan Laboratory
LinkOut - more resources
Full Text Sources

