Association of Specific Gene Mutations with Immunoglobulin Heavy-Chain Variable Region and Chromosomal Alterations in Chronic Lymphocytic Leukemia Patients in India
- PMID: 39873992
- PMCID: PMC12082404
- DOI: 10.31557/APJCP.2025.26.1.109
Association of Specific Gene Mutations with Immunoglobulin Heavy-Chain Variable Region and Chromosomal Alterations in Chronic Lymphocytic Leukemia Patients in India
Abstract
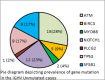
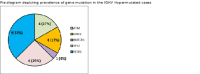
Chronic lymphocytic leukemia (CLL) is a less common hematological malignancy in Indian people. It accounts for less than 5% of all leukemias. Information on genomic alteration in CLL is limited immunoglobulin heavy-chain variable region (IGHV) mutational status is considered the most reliable prognostic marker. In this study, we performed mutation analysis of significantly mutated genes of CLL and correlated them with the IGHV mutational status and cytogenetic alterations. We included 97 patients in this study; 36 were IGHV hypermutated, and 61 were IGHV unmutated. We observed frequent mutations in TP53 (16.4%), ATM (19.5%), SF3B1 (18.5%), and NOTCH1 (14.2%). NOTCH1 mutations were significantly observed in patients with unmutated IGHV. We observed that patients with no mutations in ATM, NOTCH1, or TP53 had chromosomal alterations (del 11q, del 13q, del 17q, and trisomy 21) identified by FISH. Our results have shown mutations in essential genes and their association with IGHV status. Overall, specific gene mutations, IGHV status, and chromosomal alterations can provide information on prognosis.
Keywords: Chronic Lymphocytic Leukemia; NOTCH1; hematological malignancy.
Figures
Similar articles
-
Frequencies of SF3B1, NOTCH1, MYD88, BIRC3 and IGHV mutations and TP53 disruptions in Chinese with chronic lymphocytic leukemia: disparities with Europeans.Oncotarget. 2015 Mar 10;6(7):5426-34. doi: 10.18632/oncotarget.3101. Oncotarget. 2015. PMID: 25605254 Free PMC article.
-
Impact of gene mutations and chromosomal aberrations on progression-free survival in chronic lymphocytic leukemia patients treated with front-line chemoimmunotherapy: Clinical practice experience.Leuk Res. 2019 Jun;81:75-81. doi: 10.1016/j.leukres.2019.04.015. Epub 2019 Apr 25. Leuk Res. 2019. PMID: 31054420
-
Distinct immunoglobulin heavy chain variable region gene repertoire and lower frequency of del(11q) in Taiwanese patients with chronic lymphocytic leukaemia.Br J Haematol. 2019 Oct;187(1):82-92. doi: 10.1111/bjh.16051. Epub 2019 Jun 23. Br J Haematol. 2019. PMID: 31230372 Free PMC article.
-
Chronic lymphocytic leukemia: a clinical and molecular heterogenous disease.Cancer Genet. 2013 Mar;206(3):49-62. doi: 10.1016/j.cancergen.2013.01.003. Epub 2013 Mar 24. Cancer Genet. 2013. PMID: 23531595 Review.
-
Whole-exome and whole-genome sequencing in chronic lymphocytic leukemia: new biomarkers to target.Pharmacogenomics. 2020 Aug;21(13):957-962. doi: 10.2217/pgs-2020-0022. Epub 2020 Aug 17. Pharmacogenomics. 2020. PMID: 32799640 Review.
References
-
- Chiorazzi N, Rai KR, Ferrarini M. Chronic lymphocytic leukemia. N Engl J Med. 2005;352(8):804–15. - PubMed
-
- Hamblin TJ, Davis Z, Gardiner A, Oscier DG, Stevenson FK. Unmutated ig v(h) genes are associated with a more aggressive form of chronic lymphocytic leukemia. Blood. 1999;94(6):1848–54. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous