Construction of a rodent neural network-skeletal muscle assembloid that simulate the postnatal development of spinal cord motor neuronal network
- PMID: 39880975
- PMCID: PMC11779978
- DOI: 10.1038/s41598-025-88292-x
Construction of a rodent neural network-skeletal muscle assembloid that simulate the postnatal development of spinal cord motor neuronal network
Abstract
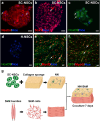
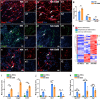
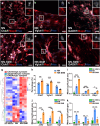
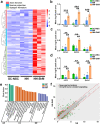
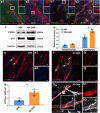
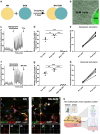
Neuromuscular diseases usually manifest as abnormalities involving motor neurons, neuromuscular junctions, and skeletal muscle (SkM) in postnatal stage. Present in vitro models of neuromuscular interactions require a long time and lack neuroglia involvement. Our study aimed to construct rodent bioengineered spinal cord neural network-skeletal muscle (NN-SkM) assembloids to elucidate the interactions between spinal cord neural stem cells (SC-NSCs) and SkM cells and their biological effects on the development and maturation of postnatal spinal cord motor neural circuits. After coculture with SkM cells, SC-NSCs developed into neural networks (NNs) and exhibited a high proportion of glutamatergic and cholinergic neurons, low proportion of neuroglia and gamma-aminobutyric acidergic neurons, and increased expression of synaptic markers. In NN-SkM assembloids, the acetylcholine receptors of SkM cells were upregulated, generating neuromuscular junction-like structures with NNs. The amplitude and frequency of SkM cell contraction in NN-SkM assembloids were increased by optogenetic and glutamate stimulation and blocked by tetrodotoxin and dizocilpine, respectively, confirming the existence of multisynaptic motor NNs. The coculture process involves the secretion of neurotrophin-3 and insulin growth factor-1 by SkM cells, which activate the related ERK-MAPK and PI3K-AKT signaling pathways in NNs. Inhibition of the ERK-MAPK and PI3K-AKT pathways significantly reduces neuronal differentiation and synaptic maturation of neural cells in NN-SkM assembloids, while also decreasing acetylcholine receptor formation on SkM cells. In brief, NN-SkM assembloids simulate the composition of spinal cord motor NNs and respond to motor regulatory signals, providing an in vitro model for studying postnatal development and maturation of spinal cord motor NNs.
Keywords: Motor neural network; Neuromuscular interaction; Skeletal muscle; Spinal cord neural stem cells; Tissue engineering.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no competing interests.
Figures

Similar articles
-
Engineering Assembloids to Mimic Graft-Host Skeletal Muscle Interaction.Adv Healthc Mater. 2025 Jul;14(17):e2404111. doi: 10.1002/adhm.202404111. Epub 2025 May 5. Adv Healthc Mater. 2025. PMID: 40320876 Free PMC article.
-
Skeletal muscle, neuromuscular organoids and assembloids: a scoping review.EBioMedicine. 2025 Aug;118:105825. doi: 10.1016/j.ebiom.2025.105825. Epub 2025 Jun 26. EBioMedicine. 2025. PMID: 40578028 Free PMC article. Review.
-
Combination of induced pluripotent stem cell-derived motor neuron progenitor cells with irradiated brain-derived neurotrophic factor over-expressing engineered mesenchymal stem cells enhanced restoration of axonal regeneration in a chronic spinal cord injury rat model.Stem Cell Res Ther. 2024 Jun 18;15(1):173. doi: 10.1186/s13287-024-03770-9. Stem Cell Res Ther. 2024. PMID: 38886817 Free PMC article.
-
Characterization of Neural Stem Cells of the Human Embryonic Fetus Across Regions of the Central Nervous System and Through Standard Gestation Period Assessments.Dev Neurobiol. 2025 Jul;85(3):e22980. doi: 10.1002/dneu.22980. Dev Neurobiol. 2025. PMID: 40495561
-
Association between neural stem/progenitor cells and biomaterials in spinal cord injury therapies: A systematic review and network meta-analysis.Acta Biomater. 2024 Jul 15;183:50-60. doi: 10.1016/j.actbio.2024.06.011. Epub 2024 Jun 12. Acta Biomater. 2024. PMID: 38871200
References
-
- Leng, Y. et al. Advances in in vitro models of neuromuscular junction: Focusing on organ-on-a-chip, organoids, and biohybrid robotics. Adv. Mater. Deerfield Beach Fla.35, e2211059 (2023). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 2024A1515012413/Natural Science Foundation of Guangdong Province, China
- 2021B1515020045/Natural Science Foundation of Guangdong Province, China
- 81891003/Chinese National Natural Science Foundation
- 2022YFA1105900/National Key Research and Development Program of China
- 82371400/National Natural Science Foundation of China
LinkOut - more resources
Full Text Sources
Miscellaneous

