The impact of signaling pathways on the desmosome ultrastructure in pemphigus
- PMID: 39882246
- PMCID: PMC11774707
- DOI: 10.3389/fimmu.2024.1497241
The impact of signaling pathways on the desmosome ultrastructure in pemphigus
Abstract
Introduction: The autoantibody-driven disease pemphigus vulgaris (PV) impairs desmosome adhesion in the epidermis. In desmosomes, the pemphigus autoantigens desmoglein 1 (Dsg1) and Dsg3 link adjacent cells. Dsgs are clustered by plaque proteins and linked to the keratin cytoskeleton by desmoplakin (Dp). The aim of this study was to identify the impact of several PV-related signaling pathways on desmosome ultrastructure.
Methods: STED microscopy, Dispase-based dissociation assay.
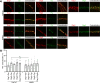
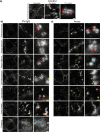
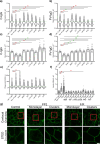
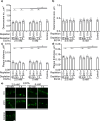
Results: As observed using STED microscopy, pemphigus autoantibodies (PV-IgG) reduced desmosome number, decreased desmosome size, increased plaque distance and thickness and caused loss of adhesion. Decreased desmosome number, increased plaque distance and thickness and loss of adhesion correlate with features found for newly assembled immature desmosomes, observed after Ca2+ depletion and repletion. This was paralleled by plaque asymmetry, keratin filament retraction and fragmentation of Dsg1 and Dsg3 immunostaining. Inhibition of each individual signaling pathway investigated here prevented the loss of adhesion and ameliorated keratin retraction. In addition, inhibition of p38MAPK or PLC completely rescued all parameters of desmosomes ultrastructure and increased desmosome number under basal conditions. In contrast, inhibition of MEK1/2 was only partially protective for desmosome size and plaque thickness, whereas inhibition of Src or increase of cAMP decreased desmosome size but increased the desmosome number even in the presence of PV-IgG.
Discussion: Alterations of the desmosomal plaque ultrastructure are closely related to loss of adhesion and regulated differently by signaling pathways involved in pemphigus pathogenesis. This insight may allow identification of novel treatment options targeting specific steps of desmosome turn-over in the future.
Keywords: adhesion; autoantibodies; autoimmune disease; desmosomes; epidermis; pemphigus; skin.
Copyright © 2025 Schmitt, Huber, Pircher, Schmidt and Waschke.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures





References
-
- Schmidt E, Goebeler M, Hertl M, Sárdy M, Sitaru C, Eming R, et al. S2k-Leitlinie zur Diagnostik des Pemphigus vulgaris/foliaceus und des bullösen Pemphigoids. JDDG: J der Deutschen Dermatologischen Gesellschaft. (2015) 13:713–27. doi: 10.1111/ddg.40_12612 - DOI
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

