Exploring the plant lipidome: techniques, challenges, and prospects
- PMID: 39883225
- PMCID: PMC11740875
- DOI: 10.1007/s44307-024-00017-9
Exploring the plant lipidome: techniques, challenges, and prospects
Abstract
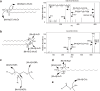
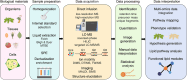
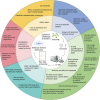
Plant lipids are a diverse group of biomolecules that play essential roles in plant architecture, physiology, and signaling. To advance our understanding of plant biology and facilitate innovations in plant-based product development, we must have precise methods for the comprehensive analysis of plant lipids. Here, we present a comprehensive overview of current research investigating plant lipids, including their structures, metabolism, and functions. We explore major lipid classes, i.e. fatty acids, glyceroglycolipids, glycerophospholipids, sphingolipids, and phytosterols, and discuss their subcellular distributions. Furthermore, we emphasize the significance of lipidomics research techniques, particularly chromatography-mass spectrometry, for accurate lipid analysis. Special attention is given to lipids as crucial signal receptors and signaling molecules that influence plant growth and responses to environmental challenges. We address research challenges in lipidomics, such as in identifying and quantifying lipids, separating isomers, and avoiding batch effects and ion suppression. Finally, we delve into the practical applications of lipidomics, including its integration with other omics methodologies, lipid visualization, and innovative analytical approaches. This review thus provides valuable insights into the field of plant lipidomics and its potential contributions to plant biology.
Keywords: Biomolecules; Environmental stress; Lipidomics; Mass spectrometry; Plant lipids.
© 2024. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Not applicable. Consent for publication: All authors approved the final manuscript and the submission to this journal. Competing interests: The authors have no competing interests to declare that are relevant to the content of this article.
Figures

References
-
- Annesley TM. Ion suppression in mass spectrometry. Clin Chem. 2003;49:1041–4. - PubMed
-
- Bartels D, Dörmann P. Plant Lipids: Methods and Protocols. Springer: Humama Press; 2021.
-
- Beale DJ, Pinu FR, Kouremenos KA, Poojary MM, Narayana VK, Boughton BA, Kanojia K, Dayalan S, Jones O, et al. Review of recent developments in GC-MS approaches to metabolomics-based research. Metabolomics. 2018;14:152. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
