Bio-nanopore technology for biomolecules detection
- PMID: 39883334
- PMCID: PMC11740844
- DOI: 10.1007/s44307-024-00051-7
Bio-nanopore technology for biomolecules detection
Abstract
Bio-nanopore technology holds great promise in biomacromolecule detection, with its high throughput and low cost positioning it as an ideal detection tool. This technology employs a unique detection mechanism that utilizes nanoscale pores to rapidly and sensitively convert biological molecules interactions into electrical signals, enabling real-time, single-molecule detection with exceptional sensitivity. This review focuses on the latest advancements in this technology across various domains, including DNA and RNA sequencing, protein detection, and small molecule identification. Additionally, future trends are explored, providing a comprehensive and in-depth perspective on the role of bio-nanopore technology in biomolecule detection.
Keywords: Bio-nanopore; DNA sequencing; Protein detection; RNA sequencing; Small molecule detection.
© 2024. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Not applicable. Consent for publication: All authors approved the final manuscript and the submission to this journal. Competing interest: The authors declare that they have no competing interests.
Figures
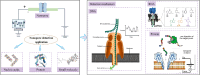



References
-
- de Cesare M, Mwenda M, Jeffreys AE, Chirwa J, Drakeley C, Schneider K, Mambwe B, Glanz K, Ntalla C, Carrasquilla M, Portugal S, Verity RJ, Bailey JA, Ghinai I, Busby GB, Hamainza B, Hawela M, Bridges DJ, Hendry JA. Flexible and cost-effective genomic surveillance of P. falciparum malaria with targeted nanopore sequencing. Nat Commun. 2024;15:1413. - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources

