A model of zymogen factor XII: insights into protease activation
- PMID: 39883942
- PMCID: PMC12018978
- DOI: 10.1182/bloodadvances.2025015842
A model of zymogen factor XII: insights into protease activation
Abstract
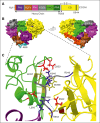
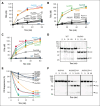
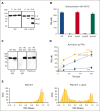
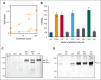
In plasma, the zymogens factor XII (FXII) and prekallikrein reciprocally convert each other to the proteases FXIIa and plasma kallikrein (PKa). PKa cleaves high-molecular-weight kininogen (HK) to release bradykinin, which contributes to regulation of blood vessel tone and permeability. Plasma FXII is normally in a "closed" conformation that limits activation by PKa. When FXII binds to a surface during contact activation it assumes an "open" conformation that increases the rate of activation by PKa. Mutations in FXII that disrupt the closed conformation have been identified in patients with conditions associated with excessive bradykinin formation. Using FXII structures from the AlphaFold database, we generated models for the closed form of human FXII that we tested with site-directed mutagenesis. The models predict multiple interactions between the fibronectin type 2 (FN2), kringle, and catalytic domains involving highly conserved amino acids that restrict access to the FXII activation cleavage sites. Based on the model, we expressed FXII with single-amino acid substitutions and studied their effects on FXII activation by PKa. Replacements for Arg36 in the FN2 domain; Glu225, Asp253, or Trp268 in the kringle domain; or Lys346 near the activation cleavage site were activated >10-fold faster by PKa than wild-type FXII. Adding these proteins to plasma resulted in rapid HK cleavage due to markedly enhanced reciprocal activation with prekallikrein. The results support a model that explains the behavior of FXII in solution. Conformational changes involving the identified amino acids likely occur when FXII binds to a surface to facilitate activation.
© 2025 American Society of Hematology. Published by Elsevier Inc. Licensed under Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International (CC BY-NC-ND 4.0), permitting only noncommercial, nonderivative use with attribution. All other rights reserved.
Conflict of interest statement
Conflict-of-interest disclosure: D.G. receives consultant fees from Anthos, Bayer, Bristol Myers Squibb, and Kanssen with an interest in inhibition of contact activation and the kallikrein-kinin system for therapeutic purposes. The remaining authors declare no competing financial interests.
Figures


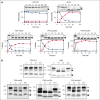
References
-
- de Maat S, Maas C. FXII: form determines function. J Thromb Haemost. 2016;14(8):1498–1506. - PubMed
-
- Maas C, Renné T. Coagulation factor XII in thrombosis and inflammation. Blood. 2018;131(17):1903–1909. - PubMed
-
- Schmaier AH. The contact activation and kallikrein/kinin systems: pathophysiologic and physiologic activities. J Thromb Haemost. 2016;14(1):28–39. - PubMed
-
- Long AT, Kenne E, Jung R, Fuchs TA, Renné T. Contact system revisited: an interface between inflammation, coagulation, and innate immunity. J Thromb Haemost. 2016;14(3):427–437. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

