Characterization of TNG348: A Selective, Allosteric USP1 Inhibitor That Synergizes with PARP Inhibitors in Tumors with Homologous Recombination Deficiency
- PMID: 39886906
- PMCID: PMC12046316
- DOI: 10.1158/1535-7163.MCT-24-0515
Characterization of TNG348: A Selective, Allosteric USP1 Inhibitor That Synergizes with PARP Inhibitors in Tumors with Homologous Recombination Deficiency
Abstract
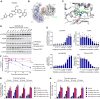
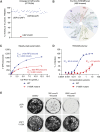
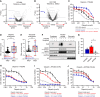
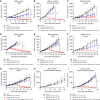
Inhibition of the deubiquitinating enzyme USP1 can induce synthetic lethality in tumors characterized by homologous recombination deficiency (HRD) and represents a novel therapeutic strategy for the treatment of BRCA1/2-mutant cancers, potentially including patients whose tumors have primary or acquired resistance to PARP inhibitors (PARPi). In this study, we present a comprehensive characterization of TNG348, an allosteric, selective, and reversible inhibitor of USP1. TNG348 induces dose-dependent accumulation of ubiquitinated protein substrates both in vitro and in vivo. CRISPR screens show that TNG348 exerts its antitumor effect by disrupting the translesion synthesis pathway of DNA damage tolerance through RAD18-dependent ubiquitinated PCNA. Although TNG348 and PARPi share the ability to selectively kill HRD tumor cells, CRISPR screens reveal that TNG348 and PARPi do so through discrete mechanisms. Particularly, knocking out PARP1 causes resistance to PARPi but sensitizes cells to TNG348 treatment. Consistent with these findings, combination of TNG348 with PARPi leads to synergistic antitumor effects in HRD tumors, resulting in tumor growth inhibition and regression in multiple mouse xenograft tumor models. Importantly, our data on human cancer models further show that the addition of TNG348 to PARPi treatment can overcome acquired PARPi resistance in vivo. Although the clinical development of TNG348 has been discontinued because of unexpected liver toxicity in patients (NCT06065059), the present data provide preclinical and mechanistic support for the continued exploration of USP1 as a drug target for the treatment of patients with BRCA1/2-mutant or HRD cancers.
©2025 The Authors; Published by the American Association for Cancer Research.
Conflict of interest statement
A. Simoneau reports other support from Tango Therapeutics outside the submitted work. C.B. Pratt reports other support from Tango Therapeutics outside the submitted work. S.S. Rajeswaran reports other support from Tango Therapeutics outside the submitted work. H.-J. Wu reports other support from Tango Therapeutics outside submitted work. C.G. Comer reports other support from Tango Therapeutics outside submitted work. S. Sudsakorn reports other support from Tango Therapeutics outside the submitted work. W. Zhang reports other support from Tango Therapeutics outside the submitted work. S. Liu reports other support from Tango Therapeutics outside submitted work. S.R. Meier reports other support from Tango Therapeutics outside the submitted work. A.H. Choi reports other support from Tango Therapeutics outside the submitted work. T. Khendu reports other support from Tango Therapeutics outside the submitted work. H. Stowe reports other support from Tango Therapeutics outside the submitted work. B. Shen reports other support from Tango Therapeutics outside the submitted work. D.A. Whittington reports other support from Tango Therapeutics and personal fees and other support from Sesame Therapeutics outside the submitted work. Y. Chen reports other support from Tango Therapeutics outside the submitted work. Y. Yu reports personal fees from Tango Therapeutics during the conduct of the study, as well as holding Tango Therapeutics stocks and options. W.D. Mallender reports other support from Tango Therapeutics outside the submitted work. T. Feng reports other support from Tango Therapeutics outside the submitted work. J.N. Andersen reports other support from Tango therapeutics during the conduct of the study. J.P. Maxwell reports other support from Tango Therapeutics outside submitted work. S. Throner reports other support from Tango Therapeutics outside the submitted work, as well as a patent for WO2022197892 pending.
Figures


Similar articles
-
The PARP1 selective inhibitor saruparib (AZD5305) elicits potent and durable antitumor activity in patient-derived BRCA1/2-associated cancer models.Genome Med. 2024 Aug 26;16(1):107. doi: 10.1186/s13073-024-01370-z. Genome Med. 2024. PMID: 39187844 Free PMC article.
-
The USP1 Inhibitor KSQ-4279 Overcomes PARP Inhibitor Resistance in Homologous Recombination-Deficient Tumors.Cancer Res. 2024 Oct 15;84(20):3419-3434. doi: 10.1158/0008-5472.CAN-24-0293. Cancer Res. 2024. PMID: 39402989 Free PMC article.
-
Single-Stranded DNA Gap Accumulation Is a Functional Biomarker for USP1 Inhibitor Sensitivity.Cancer Res. 2024 Oct 15;84(20):3435-3446. doi: 10.1158/0008-5472.CAN-23-4007. Cancer Res. 2024. PMID: 38885312 Free PMC article.
-
New Targeted Agents in Gynecologic Cancers: Synthetic Lethality, Homologous Recombination Deficiency, and PARP Inhibitors.Curr Treat Options Oncol. 2016 Mar;17(3):12. doi: 10.1007/s11864-015-0378-9. Curr Treat Options Oncol. 2016. PMID: 26931795 Free PMC article. Review.
-
Alternate therapeutic pathways for PARP inhibitors and potential mechanisms of resistance.Exp Mol Med. 2021 Jan;53(1):42-51. doi: 10.1038/s12276-021-00557-3. Epub 2021 Jan 25. Exp Mol Med. 2021. PMID: 33487630 Free PMC article. Review.
Cited by
-
USP1/UAF1 targets polyubiquitinated PCNA with an exo-cleavage mechanism that can temporarily enrich for monoubiquitinated PCNA.Nat Commun. 2025 Jul 30;16(1):6991. doi: 10.1038/s41467-025-61768-0. Nat Commun. 2025. PMID: 40739138 Free PMC article.
References
-
- de Bono J, Mateo J, Fizazi K, Saad F, Shore N, Sandhu S, et al. . Olaparib for metastatic castration-resistant prostate cancer. N Engl J Med 2020;382:2091–102. - PubMed
-
- Robson M, Im S-A, Senkus E, Xu B, Domchek SM, Masuda N, et al. . Olaparib for metastatic breast cancer in patients with a germline BRCA mutation. N Engl J Med 2017;377:523–33. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

