PPARα-mediated lipid metabolism reprogramming supports anti-EGFR therapy resistance in head and neck squamous cell carcinoma
- PMID: 39890801
- PMCID: PMC11785796
- DOI: 10.1038/s41467-025-56675-3
PPARα-mediated lipid metabolism reprogramming supports anti-EGFR therapy resistance in head and neck squamous cell carcinoma
Abstract
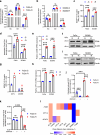
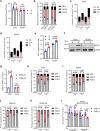
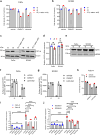
Anti-epidermal growth factor receptor (EGFR) therapy (cetuximab) shows a limited clinical benefit for patients with locally advanced or recurrent/metastatic head and neck squamous cell carcinoma (HNSCC), due to the frequent occurrence of secondary resistance mechanisms. Here we report that cetuximab-resistant HNSCC cells display a peroxisome proliferator-activated receptor alpha (PPARα)-mediated lipid metabolism reprogramming, with increased fatty acid uptake and oxidation capacities, while glycolysis is not modified. This metabolic shift makes cetuximab-resistant HNSCC cells particularly sensitive to a pharmacological inhibition of either carnitine palmitoyltransferase 1A (CPT1A) or PPARα in 3D spheroids and tumor xenografts in mice. Importantly, the PPARα-related gene signature, in human clinical datasets, correlates with lower response to anti-EGFR therapy and poor survival in HNSCC patients, thereby validating its clinical relevance. This study points out lipid metabolism rewiring as a non-genetic resistance-causing mechanism in HNSCC that may be therapeutically targeted to overcome acquired resistance to anti-EGFR therapy.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures





References
-
- Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin.71, 209–249 (2021). - PubMed
-
- Leemans, C. R., Snijders, P. J. F. & Brakenhoff, R. H. The molecular landscape of head and neck cancer. Nat. Rev. Cancer18, 269–282 (2018). - PubMed
-
- Cramer, J. D., Burtness, B., Le, Q. T. & Ferris, R. L. The changing therapeutic landscape of head and neck cancer. Nat. Rev. Clin. Oncol.16, 669–683 (2019). - PubMed
-
- Burtness, B. et al. Pembrolizumab alone or with chemotherapy versus cetuximab with chemotherapy for recurrent or metastatic squamous cell carcinoma of the head and neck (KEYNOTE-048): a randomised, open-label, phase 3 study. Lancet394, 1915–1928 (2019). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

