This is a preprint.
A Robust Proteomics-Based Method for Identifying Preferred Protein Targets of Synthetic Glycosaminoglycan Mimetics
- PMID: 39896571
- PMCID: PMC11785238
- DOI: 10.1101/2025.01.23.634492
A Robust Proteomics-Based Method for Identifying Preferred Protein Targets of Synthetic Glycosaminoglycan Mimetics
Abstract
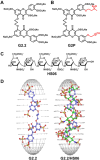
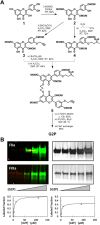
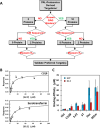
A robust technology is critically needed for identifying preferred protein targets of glycosaminoglycans (GAGs), and synthetic mimetics thereof, in biological milieu. We present a robust 10-step strategy for identification and validation of preferred protein targets of highly sulfated, synthetic, small, GAG-like molecules using diazirine-based photoaffinity labeling-proteomics approach. Our work reveals that optimally designed, homogeneous probes based on minimalistic photoactivation and affinity pulldown groups coupled with rigorous proteomics, biochemical and orthogonal validation steps offer excellent potential to identify preferred targets of GAG mimetics from the potentially numerous possible targets that cloud GAG interaction studies. Application of this 10-step strategy for a promising highly sulfated, small GAG mimetic led to identification of only a handful of preferred targets in human plasma. This new robust strategy will greatly aid drug discovery and development efforts involving GAG sequences, or sulfated small mimetics thereof, as leads.
Keywords: glycosaminoglycan-binding proteins; glycosaminoglycans; interactome; photoaffinity; proteomics.
Figures

References
-
- Perez S, Makshakova O, Angulo J, Bedini E, Bisio A, de Paz JL, Fadda E, Guerrini M, Hricovini M, Hricovini M, Lisacek F, Nieto PM, Pagel K, Paiardi G, Richter R, Samsonov SA, Vivès RR, Nikitovic D, Ricard Blum S. Glycosaminoglycans: What remains to be deciphered? JACS Au. 2023. Mar 2;3(3):628–656. - PMC - PubMed
-
- Capila I, Linhardt RJ. Heparin-protein interactions. Angew Chem Int Ed Engl. 2002. Feb 1;41(3):391–412. - PubMed
-
- Esko JD, Selleck SB. Order out of chaos: assembly of ligand binding sites in heparan sulfate. Annu Rev Biochem. 2002;71:435–71. - PubMed
-
- Vallet SD, Berthollier C, Ricard-Blum S. The glycosaminoglycan interactome 2.0. Am J Physiol Cell Physiol. 2022. Jun 1;322(6):C1271–C1278. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
