Chronic stress induces depression through MDGA1-Neuroligin2 mediated suppression of inhibitory synapses in the lateral habenula
- PMID: 39897557
- PMCID: PMC11780514
- DOI: 10.7150/thno.104282
Chronic stress induces depression through MDGA1-Neuroligin2 mediated suppression of inhibitory synapses in the lateral habenula
Abstract
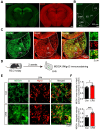
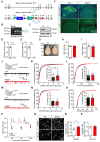
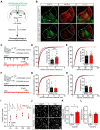
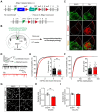
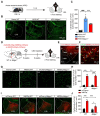
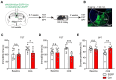
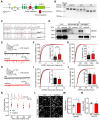
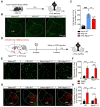
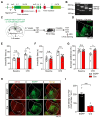
Rationale: The hyperactivity of lateral habenula (LHb) has been implicated in the pathophysiology of depression, but the regulatory mechanisms of inhibitory synapses in this context remains unclear. MDGA1 and neuroligin2 (Nlgn2), both regulators of inhibitory synapses, selectively interact in the LHb. We aimed to investigate if their interaction contributes to chronic restrained stress (CRS)-induced depression by modulating inhibitory synapses. Methods: Transgenic mouse models were established to conditional knockout/recover of MDGA1 expression or knockin Nlgn2 variant incapable of binding MDGA1 in the LHb, using viral Cre-recombinase expression. Synaptic function and density were assessed through electrophysiology and immunostaining, respectively. An acute restrained stress (ARS) model and chemogenetic activation of the lateral hypothalamus (LH) were used to stimulate the LHb. Behavioral tests related to depression were conducted following CRS. Results: MDGA1 and Nlgn2 selectively interacted in the LHb, which was elevated following CRS. Germline knockout of MDGA1 increased inhibitory transmission and GABAergic synapse density in the LHb, effects that were reversed by adult re-expression of MDGA1. Introduction of the Nlgn2 variant incapable of binding MDGA1 similarly enhanced inhibitory transmission and increased GABAergic synapse density in the LHb. Both germline MDGA1 deficiency and introduction of the Nlgn2 variant mitigated ARS- and LH activation-induced LHb neuron hyperactivation. MDGA1 deficiency in the LHb during adulthood increased inhibitory synaptic strength and conferred significant resistance to CRS-induced depressive behaviors, similar to the effects of introducing the Nlgn2 variant in the LHb. Conclusions: Our findings suggests that MDGA1-mediated suppression of Nlgn2 facilitates depression onset through limiting GABAergic synapse formation within the LHb. Targeting MDGA1/Nlgn2 complexes residing at GABAergic synapses within the lateral habenula may be viable for alleviating core behavioral symptoms of major depression.
Keywords: Depression; Inhibitory synapses; Lateral habenula; MDGA1; Nlgn2; Stress.
© The author(s).
Conflict of interest statement
Competing Interests: The authors have declared that no competing interest exists.
Figures
References
-
- Mazure CM, Maciejewski PK. A model of risk for major depression: effects of life stress and cognitive style vary by age. Depress Anxiety. 2003;17:26–33. - PubMed
-
- Willner P, Scheel-Krüger J, Belzung C. The neurobiology of depression and antidepressant action. Neurosci Biobehav Rev. 2013;37:2331–71. - PubMed
-
- Berton O, Hahn CG, Thase ME. Are we getting closer to valid translational models for major depression? Science. 2012;338:75–9. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

