Interspecies signaling modulates the biosynthesis of antimicrobial secondary metabolites related to biological control activities of Pseudomonas fluorescens 2P24
- PMID: 39898669
- PMCID: PMC11878095
- DOI: 10.1128/spectrum.01886-24
Interspecies signaling modulates the biosynthesis of antimicrobial secondary metabolites related to biological control activities of Pseudomonas fluorescens 2P24
Abstract
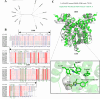
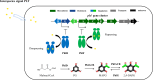
Signaling between rhizosphere microorganisms is crucial in bacteria interaction and communication, shaping the rhizomicrobiome. Plant growth-promoting bacterium Pseudomonas produces a spectrum of important antibiotics to inhibit plant pathogens, albeit with an associated metabolic burden. Antibiotics could function as intra- and interspecies signals at subinhibitory concentrations to coordinate gene expression and microbial behaviors. In this work, we characterized pyoluteorin as an interspecies signal that modulates the biosynthesis of 2,4-diacetylphloroglucinol (2,4-DAPG), a broad-spectrum biocontrol agent, in non-pyoluteorin-producing Pseudomonas fluorescens 2P24. We demonstrated that the key transcriptional repressor PhlF from the 2,4-DAPG biosynthetic gene cluster spontaneously senses pyoluteorin, enhancing repression of the phlA promoter activity and inhibiting 2,4-DAPG synthesis in P. fluorescens 2P24. Pyoluteorin also binds to another transcriptional repressor, PhlH, from the 2,4-DAPG biosynthetic gene cluster, subsequently releasing the transcription of phlG, which facilitates the hydrolysis of 2,4-DAPG. Both PhlF and PhlH are simultaneously involved in sensing exogenous pyoluteorin to regulate the 2,4-DAPG biosynthetic operon, playing a crucial role in controlling antibiotic metabolites in response to environmental changes. Further phylogenetic and structural analyses demonstrated that PhlH and PhlF are widely distributed across Pseudomonas spp. with conserved ligand-binding domains. The findings shed new light on the regulatory mechanism of 2,4-DAPG biosynthesis underlying interspecies signaling by pyoluteorin and provide invaluable clues for the rational design of co-inhabiting Pseudomonas spp. as biocontrol agents.
Importance: Rhizosphere microorganisms release vital signals that shape microbial communities, with antibiotics at low concentrations acting as intra- and interspecies signals. However, the mechanisms of these signals in coordinating gene expression are unclear. In non-pyoluteorin-producing Pseudomonas fluorescens 2P24, pyoluteorin was identified as an interspecies signal that regulates the phl biosynthesis gene cluster for 2,4-DAPG production. TetR family repressors PhlH and PhlF were found to positively regulate 2,4-DAPG hydrolysis and negatively regulate its synthesis in response to pyoluteorin. Structural modeling and docking analyses revealed the interactions between pyoluteorin and both PhlH and PhlF, modulating gene expression. Phylogenetic analyses showed a wide distribution of PhlH and PhlF across Pseudomonas spp. with conserved ligand-binding domains. These findings deepen our understanding of interspecies signaling mechanisms and highlight the potential for designing co-inhabiting Pseudomonas spp. as effective biocontrol agents.
Keywords: 2,4-diacetylphloroglucinol; Pseudomonas; interspecies signaling; pyoluteorin; transcriptional repressor.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Pyoluteorin regulates the biosynthesis of 2,4-DAPG through the TetR family transcription factor PhlH in Pseudomonas protegens Pf-5.Appl Environ Microbiol. 2024 Apr 17;90(4):e0174323. doi: 10.1128/aem.01743-23. Epub 2024 Mar 12. Appl Environ Microbiol. 2024. PMID: 38470180 Free PMC article.
-
Transcriptional Regulator PhlH Modulates 2,4-Diacetylphloroglucinol Biosynthesis in Response to the Biosynthetic Intermediate and End Product.Appl Environ Microbiol. 2017 Oct 17;83(21):e01419-17. doi: 10.1128/AEM.01419-17. Print 2017 Nov 1. Appl Environ Microbiol. 2017. PMID: 28821548 Free PMC article.
-
Autoinduction of 2,4-diacetylphloroglucinol biosynthesis in the biocontrol agent Pseudomonas fluorescens CHA0 and repression by the bacterial metabolites salicylate and pyoluteorin.J Bacteriol. 2000 Mar;182(5):1215-25. doi: 10.1128/JB.182.5.1215-1225.2000. J Bacteriol. 2000. PMID: 10671440 Free PMC article.
-
Role of 2,4-diacetylphloroglucinol-producing fluorescent Pseudomonas spp. in the defense of plant roots.Plant Biol (Stuttg). 2007 Jan;9(1):4-20. doi: 10.1055/s-2006-924473. Epub 2006 Oct 23. Plant Biol (Stuttg). 2007. PMID: 17058178 Review.
-
Phloroglucinol Derivatives in Plant-Beneficial Pseudomonas spp.: Biosynthesis, Regulation, and Functions.Metabolites. 2021 Mar 20;11(3):182. doi: 10.3390/metabo11030182. Metabolites. 2021. PMID: 33804595 Free PMC article. Review.
References
-
- Zhalnina K, Louie KB, Hao Z, Mansoori N, da Rocha UN, Shi S, Cho H, Karaoz U, Loqué D, Bowen BP, Firestone MK, Northen TR, Brodie EL. 2018. Dynamic root exudate chemistry and microbial substrate preferences drive patterns in rhizosphere microbial community assembly. Nat Microbiol 3:470–480. doi:10.1038/s41564-018-0129-3 - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

