Plant-nanoparticles enhance anti-PD-L1 efficacy by shaping human commensal microbiota metabolites
- PMID: 39900923
- PMCID: PMC11790884
- DOI: 10.1038/s41467-025-56498-2
Plant-nanoparticles enhance anti-PD-L1 efficacy by shaping human commensal microbiota metabolites
Abstract
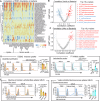
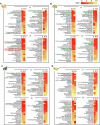
Diet has emerged as a key impact factor for gut microbiota function. However, the complexity of dietary components makes it difficult to predict specific outcomes. Here we investigate the impact of plant-derived nanoparticles (PNP) on gut microbiota and metabolites in context of cancer immunotherapy with the humanized gnotobiotic mouse model. Specifically, we show that ginger-derived exosome-like nanoparticle (GELN) preferentially taken up by Lachnospiraceae and Lactobacillaceae mediated by digalactosyldiacylglycerol (DGDG) and glycine, respectively. We further demonstrate that GELN aly-miR159a-3p enhances anti-PD-L1 therapy in melanoma by inhibiting the expression of recipient bacterial phospholipase C (PLC) and increases the accumulation of docosahexaenoic acid (DHA). An increased level of circulating DHA inhibits PD-L1 expression in tumor cells by binding the PD-L1 promoter and subsequently prevents c-myc-initiated transcription of PD-L1. Colonization of germ-free male mice with gut bacteria from anti-PD-L1 non-responding patients supplemented with DHA enhances the efficacy of anti-PD-L1 therapy compared to controls. Our findings reveal a previously unknown mechanistic impact of PNP on human tumor immunotherapy by modulating gut bacterial metabolic pathways.
© 2025. This is a U.S. Government work and not under copyright protection in the US; foreign copyright protection may apply.
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures








References
-
- Anusha, R. & Priya, S. Dietary exosome-like nanoparticles: An updated review on their pharmacological and drug delivery applications. Mol. Nutr. Food Res.66, e2200142 (2022). - PubMed
MeSH terms
Substances
Grants and funding
- IK6BX004199/Biomedical Laboratory Research and Development, VA Office of Research and Development (VA Biomedical Laboratory Research and Development)
- P30 ES030283/ES/NIEHS NIH HHS/United States
- P20 GM113226/GM/NIGMS NIH HHS/United States
- IK6 BX004199/BX/BLRD VA/United States
- P50 AA024337/AA/NIAAA NIH HHS/United States
- P20 GM125504/GM/NIGMS NIH HHS/United States
- BX005254/Biomedical Laboratory Research and Development, VA Office of Research and Development (VA Biomedical Laboratory Research and Development)
- P50-AA024337/U.S. Department of Health & Human Services | NIH | National Institute on Alcohol Abuse and Alcoholism (NIAAA)
- R01 AT008617/AT/NCCIH NIH HHS/United States
- R01AT008617/U.S. Department of Health & Human Services | NIH | National Center for Complementary and Integrative Health (NCCIH)
- P20GM125504/U.S. Department of Health & Human Services | NIH | National Institute of General Medical Sciences (NIGMS)
- P20 GM103436/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

