The genome sequence of a sawfly, Abia candens Konow, 1887
- PMID: 39912117
- PMCID: PMC11795027
- DOI: 10.12688/wellcomeopenres.23449.1
The genome sequence of a sawfly, Abia candens Konow, 1887
Abstract
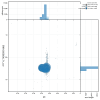
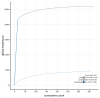
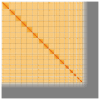
We present a genome assembly from an individual male specimen of Abia candens (sawfly; Arthropoda; Insecta; Hymenoptera; Cimbicidae). The genome sequence has a total length of 261.00 megabases. Most of the assembly (82.7%) is scaffolded into 16 chromosomal pseudomolecules. The mitochondrial genome has also been assembled and is 19.72 kilobases in length.
Keywords: Abia candens; Hymenoptera; chromosomal; genome sequence; sawfly.
Copyright: © 2025 Cunningham A et al.
Conflict of interest statement
No competing interests were disclosed.
Figures

References
-
- Bates A, Clayton-Lucey I, Howard C: Sanger Tree of Life HMW DNA fragmentation: diagenode Megaruptor ®3 for LI PacBio. protocols.io. 2023. 10.17504/protocols.io.81wgbxzq3lpk/v1 - DOI
Grants and funding
LinkOut - more resources
Full Text Sources

