Nonstop mutations cause loss of renal tumor suppressor proteins VHL and BAP1 and affect multiple stages of protein translation
- PMID: 39937911
- PMCID: PMC11817944
- DOI: 10.1126/sciadv.adr6375
Nonstop mutations cause loss of renal tumor suppressor proteins VHL and BAP1 and affect multiple stages of protein translation
Abstract
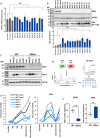
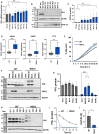
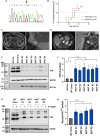
Nonstop extension or stop-loss mutations lead to the extension of a protein at its carboxyl terminus. Recently, nonstop mutations in the tumor suppressor SMAD Family Member 4 (SMAD4) have been discovered to lead to proteasomal SMAD4 degradation. However, this mutation type has not been studied in other cancer genes. Here, we explore somatic nonstop mutations in the tumor suppressor genes BRCA1 Associated Protein 1 (BAP1) and Von Hippel-Lindau (VHL) enriched in renal cell carcinoma. For BAP1, nonstop mutations generate an extremely long extension. Instead of proteasomal degradation, the extension decreases translation and depletes BAP1 messenger RNA from heavy polysomes. For VHL, the short extension leads to proteasomal degradation. Unexpectedly, the mutation alters the selection of the translational start site shifting VHL isoforms. We identify germline VHL nonstop mutations in patients leading to the early onset of severe disease manifestations. In summary, nonstop extension mutations inhibit the expression of renal tumor suppressor genes with pleiotropic effects on translation and protein stability.
Figures


References
-
- Frischmeyer P. A., van Hoof A., O’Donnell K., Guerrerio A. L., Parker R., Dietz H. C., An mRNA surveillance mechanism that eliminates transcripts lacking termination codons. Science 295, 2258–2261 (2002). - PubMed
-
- Yordanova M. M., Loughran G., Zhdanov A. V., Mariotti M., Kiniry S. J., O’Connor P. B. F., Andreev D. E., Tzani I., Saffert P., Michel A. M., Gladyshev V. N., Papkovsky D. B., Atkins J. F., Baranov P. V., AMD1 mRNA employs ribosome stalling as a mechanism for molecular memory formation. Nature 553, 356–360 (2018). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

