Genomic and Transcriptomic Analysis of the Polyploidy Cyst Nematode, Heterodera trifolii, and Heterodera schachtii
- PMID: 39940717
- PMCID: PMC11817004
- DOI: 10.3390/ijms26030948
Genomic and Transcriptomic Analysis of the Polyploidy Cyst Nematode, Heterodera trifolii, and Heterodera schachtii
Abstract
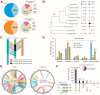
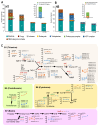
Cyst nematodes remain a major threat to global agricultural production, causing huge losses. To understand the parasitism of the cyst nematodes Heterodera trifolii (HT) and Heterodera schachtii (HS), we constructed whole-genome assemblies using short- and long-read sequencing technologies. The nematode genomes were 379 Mb and 183 Mb in size, with the integrated gene models predicting 40,186 and 18,227 genes in HT and HS, respectively. We found more than half of the genes predicted in HT (64.7%) and HS (53.2%) were collinear to their nearest neighbor H. glycines (HG). Large-scale duplication patterns in HT and segmental duplications of more than half of the orthologous genes indicate that the genome of HT is polyploid in nature. Functional analysis of the genes indicated that 65.6% of the HG genes existed within the HT genome. Most abundant genes in HT and HS were involved in gene regulation, DNA integration, and chemotaxis. Differentially expressed genes showed upregulation of cuticle structural constituent genes during egg and female stages and cytoskeletal motor activity-related genes in juvenile stage 2 (J2). Horizontal gene transfer analyses identified four new vitamin biosynthesis genes, pdxK, pdxH, pdxS, and fabG, of bacterial origin, to be first reported in HT and HS. Mitogenomes of HT, HS, and HG showed similar structure, composition, and codon usage. However, rates of substitution of bases in the gene nad4l were significantly different between HT and HS. The described genomes, transcriptomes, and mitogenomes of plant-parasitic nematodes HT and HS are potential bio-resources used to identify several strategies of control of the nematode.
Keywords: Heterodera; cyst nematode; genome assembly; transcriptome.
Conflict of interest statement
Authors Daegwan Kim and Yeisoo Yu were employed by the company DNACARE Co., Ltd. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




References
-
- van Steenbrugge J.J.M., van den Elsen S., Holterman M., Lozano-Torres J.L., Putker V., Thorpe P., Goverse A., Sterken M.G., Smant G., Helder J. Comparative genomics among cyst nematodes reveals distinct evolutionary histories among effector families and an irregular distribution of effector-associated promoter motifs. Mol. Ecol. 2023;32:1515–1529. doi: 10.1111/mec.16505. - DOI - PMC - PubMed
-
- Nicol J.M., Turner S.J., Coyne D.L., den Nijs L., Hockland S., Maafi Z.T. Current Nematode Threats to World Agriculture. In: Jones J., Gheysen G., Fenoll C., editors. Genomics and Molecular Genetics of Plant-Nematode Interactions. Springer; Dordrecht, The Nederlands: 2011. pp. 21–43. - DOI
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

