OsMAPKKK5 affects brassinosteroid signal transduction via phosphorylating OsBSK1-1 and regulates rice plant architecture and yield
- PMID: 39967024
- PMCID: PMC12018843
- DOI: 10.1111/pbi.70008
OsMAPKKK5 affects brassinosteroid signal transduction via phosphorylating OsBSK1-1 and regulates rice plant architecture and yield
Abstract
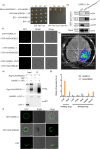
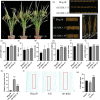
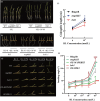
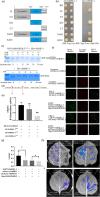
Improving plant architecture and increasing yields are the main goals of rice breeders. However, yield is a complex trait influenced by many yield-related traits. Identifying and characterizing important genes in the coordinated network regulating complex rice traits and their interactions is conducive to cultivating high-yielding rice varieties. In this study, we determined that the interaction between mitogen-activated protein kinase kinase kinase5 (OsMAPKKK5) and brassinosteroid-signalling kinase1-1 (OsBSK1-1) regulates yield-related traits in rice. Specifically, OsMAPKKK5 phosphorylates OsBSK1-1, which enhances the interaction between these two proteins, but adversely affects the OsBSK1-1-OsBRI1 (BR insensitive1) and OsBSK1-1-OsPPKL1 (protein phosphatase with two Kelch-like domains) interactions. Additionally, OsMAPKKK5 disrupts brassinosteroid signal transduction, which prevents OsBZR1 (brassinazole-resistant1) from efficiently entering the nucleus, thereby negatively modulating its function as a transcription factor regulating downstream effector genes, ultimately adversely affecting plant architecture and yield. This study revealed the relationship between the MAPK cascade and the regulatory effects of brassinosteroid on the rice grain yield involves OsMAPKKK5 and OsBSK1-1. The study data may be important for future investigations on the rice yield-regulating molecular network.
Keywords: brassinosteroids; crosstalk; mitogen‐activated protein kinase kinase kinase; plant architecture; rice yield.
© 2025 The Author(s). Plant Biotechnology Journal published by Society for Experimental Biology and The Association of Applied Biologists and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no competing financial interests.
Figures

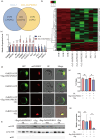
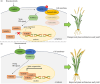
References
-
- Chen, J. , Wang, L. , Yang, Z. , Liu, H. , Chu, C. , Zhang, Z. , Zhang, Q. et al. (2021) The rice Raf‐like MAPKKK OsILA1 confers broad‐spectrum resistance to bacterial blight by suppressing the OsMAPKK4‐OsMAPK6 cascade. J. Integr. Plant Biol. 63(10), 1815–1842. - PubMed
-
- Du, H. , Yong, R. , Zhang, J. , Cai, G. , Wang, R. , Li, J. , Wang, Y. et al. (2023) OsBAK2/OsSERK2 expression is repressed by OsBZR1 to modulate brassinosteroid response and grain length in rice. J. Exp. Bot. 74(17), 4978–4993. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

