Prevalence and Virulence Profiles of Klebsiella pneumoniae Isolated From Different Animals
- PMID: 39969166
- PMCID: PMC11837280
- DOI: 10.1002/vms3.70243
Prevalence and Virulence Profiles of Klebsiella pneumoniae Isolated From Different Animals
Abstract
Background: Klebsiella pneumoniae liver abscess (KLA) is an invasive disease, and the occurrence of infection is related to its virulence factors and colonization of the host's gastrointestinal (GI) tract. Some animal-sourced isolates share virulence factors with human pathogens. However, the potential of K. pneumoniae as a zoonotic agent has not been confirmed in murine infection model.
Objectives: To identify the prevalence and virulence profiles of K. pneumoniae colonization in companion and wild animals and subsequently determine the pathogenicity of selected strains.
Methods: Forty-five K. pneumoniae isolates (45/302) were obtained from faeces of companion or wild animals. Virulence factors, gyrA polymerase chain reaction with the restriction fragment length polymorphism (PCR-RFLP) and pulsed field gel electrophoresis (PFGE) were detected and compared with our previous collection of 60 human pathogens. For KLA model and cytotoxicity test, three animal-sourced isolates, CHKP0009 (snake, K1, KpII), CHKP0021 (turtle, K2, pLVPK, KpI, cluster I) and CHKP1027 (dog, non-K1/K2, HV, KpI, cluster III), with similar genotype and/or phenotype to human pathogens were selected and evaluated for their virulence with human hypervirulent K. pneumoniae (hvKp) CG43S.
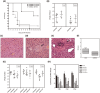
Results: The prevalence of K. pneumoniae was higher in companion than wild animals. K. pneumoniae was primarily isolated from dogs, turtles and snakes. Some animal-sourced isolates carried virulence factors and revealed phylogenetic relatedness with human pathogens. In KLA model, BALB/c mice infected with snake isolate CHKP0009 and dog isolate CHKP1027 survived for 14 days but showed significant bacterial loads in the liver and spleen. Notably, the pet turtle isolate CHKP0021 presented comparable virulence with human hvKp CG43S and induced liver abscess formation. All three selected animal-sourced isolates could colonize in the GI tract and possess cytotoxic ability. These findings demonstrated pathogenicity of the animal K. pneumoniae isolates. In addition, the high prevalence of K. pneumoniae in companion animals and some isolates with virulence profiles suggested animal-sourced K. pneumoniae has the zoonotic potential to cause human disease.
Conclusion: Animals are the natural hosts of zoonotic pathogens. Some animal-sourced K. pneumoniae isolates are not only pathogenic in vivo but also exhibit phylogenetic relatedness to human pathogens, suggesting the existence of a zoonotic risk for K. pneumoniae between these two populations.
Keywords: Hypervirulent; Klebsiella pneumoniae; liver abscess; zoonotic pathogen.
© 2025 The Author(s). Veterinary Medicine and Science published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

References
-
- Athamna, A. , Ofek I., Keisari Y., Markowitz S., Dutton G. G., and Sharon N.. 1991. “Lectinophagocytosis of Encapsulated Klebsiella pneumoniae Mediated by Surface Lectins of Guinea Pig Alveolar Macrophages and Human Monocyte‐Derived Macrophages.” Infection and Immunity 59, no. 5: 1673–1682. 10.1128/iai.59.5.1673-1682.1991. - DOI - PMC - PubMed
-
- Brendecke, J. , Homeier‐Bachmann T., Schmitz Ornés A., et al. 2022. “Multidrug‐Resistant High‐Risk Escherichia coli and Klebsiella pneumoniae Clonal Lineages Occur in Black‐Headed Gulls From Two Conservation Islands in Germany.” Antibiotics 11, no. 10: 1357. 10.3390/antibiotics11101357. - DOI - PMC - PubMed
-
- Brisse, S. , van Himbergen T., Kusters K., and Verhoef J.. 2004. “Development of a Rapid Identification Method for Klebsiella pneumoniae Phylogenetic Groups and Analysis of 420 Clinical Isolates.” Clinical Microbiology and Infection 10, no. 10: 942–945. 10.1111/j.1469-0691.2004.00973.x. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

