Cardiomyocyte-restricted MIAT deletion is sufficient to protect against murine myocardial infarction
- PMID: 39979325
- PMCID: PMC11842840
- DOI: 10.1038/s41420-025-02352-9
Cardiomyocyte-restricted MIAT deletion is sufficient to protect against murine myocardial infarction
Abstract
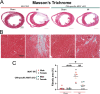
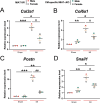
Myocardial infarction-associated transcript (MIAT), an intergenic long noncoding RNA (lncRNA), is conserved between rodents and humans and is directly linked to maladaptive cardiac remodeling in both patients and mouse models with various forms of heart failure (HF). We previously reported attenuation of cardiac stress, apoptosis, and fibrosis in a murine model of myocardial infarction (MI) with global MIAT ablation. Our transcriptomic profiling and mechanistic studies further revealed MIAT-induced activation of maladaptive genes, such as Hoxa4, Fmo2, Lrrn4, Marveld3, and Fat4. However, the source of MIAT and its contribution to MI and HF remain unknown. In this study, we generate a novel cardiomyocyte (CM)-specific MIAT conditional knockout mouse model, which exhibits improved cardiac function after MI. We further report that CM-specific MIAT ablation is sufficient to reduce cardiac damage, apoptosis, and fibrosis following chronic MI. Mechanistically, CM-specific MIAT deletion in mice leads to decreased expression of proapoptotic and pathological profibrotic genes, such as p53, Bak1, Col3a1, Col6a1, Postn, and Snail1 after chronic MI. These results enable us to begin to dissect cell-specific contributions to MIAT signaling and bolster the idea that MIAT plays a direct pathological role in CMs after MI.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures





References
-
- Takemura G, Fujiwara H. Role of apoptosis in remodeling after myocardial infarction. Pharm Ther. 2004;104:1–16. 10.1016/j.pharmthera.2004.07.005. - PubMed
-
- Thum T, Condorelli G. Long Noncoding RNAs and MicroRNAs in cardiovascular pathophysiology. Circ Res. 2015;116:751–62. 10.1161/Circresaha.116.303549. - PubMed
Grants and funding
- HT9425-24-1-0209/U.S. Department of Defense (United States Department of Defense)
- R01HL146481/U.S. Department of Health & Human Services | National Institutes of Health (NIH)
- R01 HL146481/HL/NHLBI NIH HHS/United States
- R01 HL160941/HL/NHLBI NIH HHS/United States
- 931621/AHA/American Heart Association-American Stroke Association/United States
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

