Development of a cell-penetrating peptide-based nanocomplex for long-term delivery of intact mitochondrial DNA into epithelial cells
- PMID: 39991470
- PMCID: PMC11847061
- DOI: 10.1016/j.omtn.2025.102449
Development of a cell-penetrating peptide-based nanocomplex for long-term delivery of intact mitochondrial DNA into epithelial cells
Abstract
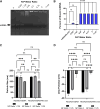
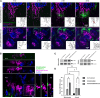
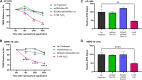
Gene therapy approaches for mitochondrial DNA (mtDNA)-associated damage/diseases have thus far been limited, and despite advancements in single gene therapy for mtDNA mutations and progress in mitochondrial transplantation, no method exists for restoring the entire mtDNA molecule in a clinically translatable manner. Here, we present for the first time a strategy to deliver an exogenous, fully intact, and healthy mtDNA template into cells to correct endogenous mtDNA mutations and deletions, with the potential to be developed into an efficient pan-therapy for inherited and/or acquired mtDNA disorders. More specifically, the novel therapeutic nanoparticle complex used in our study was generated by combining a cell-penetrating peptide (CPP) with purified mtDNA, in conjunction with a mitochondrial targeting reagent. The generated nanoparticle complexes were found to be taken up by cells and localized to mitochondria, with exogenous mtDNA retention/maintenance, along with mitochondrial RNA and protein production, observed in mitochondria-depleted ARPE-19 cells at least 4 weeks following a single treatment. These data demonstrate the feasibility of restoring mtDNA in cells via a CPP carrier, with the therapeutic potential to correct mtDNA damage independent of the number of gene mutations found within the mtDNA.
Keywords: MT: Delivery Strategies; age-related diseases; cell-penetrating peptide; mitochondria; mitochondrial DNA; mitochondrial transplantation; mtDNA; mtDNA gene therapy; mtDNA mutations; nucleic acid delivery.
Published by Elsevier Inc. on behalf of The American Society of Gene and Cell Therapy.
Conflict of interest statement
K.W., C.H., A.J., and B.R. have filed a patent application for the utilization of CPP-based mtDNA-containing nanocomplexes in the study of mitochondrial biology/metabolism/genetics and treatment of mitochondrial-related diseases.
Figures




References
-
- Kaarniranta K., Uusitalo H., Blasiak J., Felszeghy S., Kannan R., Kauppinen A., Salminen A., Sinha D., Ferrington D. Mechanisms of mitochondrial dysfunction and their impact on age-related macular degeneration. Prog. Retin. Eye Res. 2020;79:100858. doi: 10.1016/j.preteyeres.2020.100858. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

