Virulence and Genetic Diversity of Puccinia spp., Causal Agents of Rust on Switchgrass (Panicum virgatum L.) in the USA
- PMID: 40005569
- PMCID: PMC11858125
- DOI: 10.3390/pathogens14020194
Virulence and Genetic Diversity of Puccinia spp., Causal Agents of Rust on Switchgrass (Panicum virgatum L.) in the USA
Abstract
Switchgrass (Panicum virgatum L.) is an important cellulosic biofuel grass native to North America. Rust, caused by Puccinia spp. is the most predominant disease of switchgrass and has the potential to impact biomass conversion. In this study, virulence patterns were determined on a set of 38 switchgrass genotypes for 14 single-spore rust isolates from 14 field samples collected in seven states. Single nucleotide polymorphism (SNP) variation was also assessed in 720 sequenced cloned amplicons representing 654 base pairs of the elongation factor 1-α gene from the field samples. Five major haplotypes were identified differing by 11 out of the 39 SNP positions identified. STRUCTURE, Principal Coordinate Analysis, and phylogenetic analyses divided the rust population into two genetic clusters. Virginia and Georgia had the highest and lowest rust genetic diversity, respectively. Only nine accessions showed a differential disease response between the 14 isolates, allowing the identification of eight races, differing by 1-3 virulence factors. Overall, the results suggested clonal reproduction of the pathogen and a North-South differentiation via local adaptation. However, similar haplotypes and races were also recovered from several states, suggesting migration events, and highlighting the need to further investigate the switchgrass rust population structure and evolution in the USA.
Keywords: genetic diversity; rust; switchgrass; virulence.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures



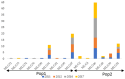

References
-
- Zhang Y., Zalapa J.E., Jakubowski A.R., Price D.L., Acharya A., Wei Y., Brummer E.C., Kaeppler S.M., Casler M.D. Post-Glacial Evolution of Panicum Virgatum: Centers of Diversity and Gene Pools Revealed by SSR Markers and CpDNA Sequences. Genetica. 2011;139:933–948. doi: 10.1007/s10709-011-9597-6. - DOI - PubMed
-
- Wright L., Turhollow A. Switchgrass Selection as a “Model” Bioenergy Crop: A History of the Process. Biomass Bioenergy. 2010;34:851–868. doi: 10.1016/j.biombioe.2010.01.030. - DOI
MeSH terms
Grants and funding
- 2015-67009-23962/Agriculture and Food Research Initiative from the USDA National Institute of Food and Agri-culture
- DE-AC05-00OR22725/Center for Bioenergy Innovation (CBI), a U.S. Department of Energy Bioenergy Research Center supported by the Office of Biological and Environmental Research in the Department of Energy (DOE) Office of Science
LinkOut - more resources
Full Text Sources
Miscellaneous

