Prevalence of Aphid-Transmitted Potyviruses in Pumpkin and Winter Squash in Georgia, USA
- PMID: 40006988
- PMCID: PMC11860210
- DOI: 10.3390/v17020233
Prevalence of Aphid-Transmitted Potyviruses in Pumpkin and Winter Squash in Georgia, USA
Abstract
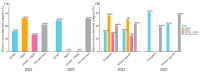
Viruses are a major pathogen challenging the sustainable production of cucurbits worldwide. Pumpkin and winter squash showed severe virus-like symptoms during the fall of 2022 and 2023 in Georgia, USA. Symptomatic leaves were collected from the field and processed for small RNA sequencing for virus identification using high-throughput sequencing (HTS). HTS analysis revealed the presence of two aphid-transmitted viruses (ATVs), zucchini yellow mosaic virus (ZYMV) and papaya ringspot virus (PRSV), along with three whitefly-transmitted viruses, cucurbit chlorotic yellows virus, cucurbit yellow stunting disorder virus, and cucurbit leaf crumple virus. The results of our study suggest a significant shift in ATV's abundance in these two crops between 2022 and 2023. According to the qPCR data in the fall of 2022, pumpkins experience an incidence of 56.25% and 31.25% of PRSV and ZYMV, respectively. Similarly, winter squash shows an incidence of 50% and 32.14% of PRSV and ZYMV, respectively. Mixed infection of both viruses was also observed in these two crops. In 2023, we observed a predominance of ZYMV in pumpkin and winter squash (61.25% and 42.50%, respectively). However, PRSV was not detected in pumpkins, and it was detected at a negligible level (0.62%) in winter squash using qPCR. Phylogenetic analysis of ZYMV-encoded coat protein (CP) and helper component-protease (HC-Pro) from Georgia suggests a close relationship with the European isolates. Conversely, PRSV-encoded CP and NIa-VPg show a more diverse evolutionary history. Overall, this research will provide valuable insights into the dynamics of ZYMV and PRSV in pumpkin and winter squash crops within the southeastern United States.
Keywords: aphids; cucurbit; high-throughput sequencing; papaya ringspot virus; prevalence; pumpkin; winter squash; zucchini yellow mosaic virus.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures



Similar articles
-
High Throughput Sequencing-Aided Survey Reveals Widespread Mixed Infections of Whitefly-Transmitted Viruses in Cucurbits in Georgia, USA.Viruses. 2021 May 26;13(6):988. doi: 10.3390/v13060988. Viruses. 2021. PMID: 34073397 Free PMC article.
-
First Report of Zucchini yellow mosaic virus in Cucurbits in Ivory Coast.Plant Dis. 2010 Nov;94(11):1378. doi: 10.1094/PDIS-06-10-0416. Plant Dis. 2010. PMID: 30743639
-
Persistent, and Asymptomatic Viral Infections and Whitefly-Transmitted Viruses Impacting Cantaloupe and Watermelon in Georgia, USA.Viruses. 2022 Jun 15;14(6):1310. doi: 10.3390/v14061310. Viruses. 2022. PMID: 35746780 Free PMC article.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2021 Apr 19;4(4):CD011535. doi: 10.1002/14651858.CD011535.pub4. Cochrane Database Syst Rev. 2021. Update in: Cochrane Database Syst Rev. 2022 May 23;5:CD011535. doi: 10.1002/14651858.CD011535.pub5. PMID: 33871055 Free PMC article. Updated.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2017 Dec 22;12(12):CD011535. doi: 10.1002/14651858.CD011535.pub2. Cochrane Database Syst Rev. 2017. Update in: Cochrane Database Syst Rev. 2020 Jan 9;1:CD011535. doi: 10.1002/14651858.CD011535.pub3. PMID: 29271481 Free PMC article. Updated.
References
-
- Food and Agriculture Organization (FAO) Global Production of Vegetables in 2022, by Type (in Million Metric Tons), in Statista. [(accessed on 15 February 2024)]. Available online: https://www.statista.com/statistics/264065/global-production-of-vegetabl...
-
- United States Department of Agriculture (USDA) Vegetables 2023 Summary. United States Department of Agriculture, National Agricultural Statistics Service; Washington, DC, USA: 2024. [(accessed on 7 January 2025)]. pp. 6–84. Available online: https://downloads.usda.library.cornell.edu/usda-esmis/files/02870v86p/qz....
-
- Evranuz E.O., Arduzlar-Kagan D. Winter squash and pumpkins. In: Hui Y.H., Ozgul Evranuz E., editors. Handbook of Vegetable Preservation and Processing. 2nd ed. CRC Press; Boca Raton, FL, USA: 2015. pp. 671–690.
-
- Kates H.R. Pumpkins, squashes, and gourds (Cucurbita L.) of North America. In: Greene S.L., Williams K.A., Khoury C.K., Kantar M.B., Marek L.F., editors. North American Crop Wild Relatives. Springer; Cham, Switzerland: 2019. pp. 195–224.
Publication types
MeSH terms
Supplementary concepts
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

