Comparing Alzheimer's genes in African, European, and Amerindian induced pluripotent stem cell-derived microglia
- PMID: 40008916
- PMCID: PMC11863361
- DOI: 10.1002/alz.70031
Comparing Alzheimer's genes in African, European, and Amerindian induced pluripotent stem cell-derived microglia
Abstract
Introduction: Genome-wide association studies (GWAS) studies in Alzheimer's disease (AD) demonstrate ancestry-specific loci. Previous studies in the regulatory architecture have only been conducted in Europeans (EUs), thus studies in additional ancestries are needed. Given the prevalence of AD genes expressed in microglia, we initiated our studies in induced pluripotent stem cell (iPSC) -derived microglia.
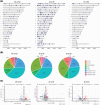
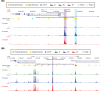
Methods: We created iPSC-derived microglia from 13 individuals of either high Amerindian (AI), African (AF), or EU global ancestry, including both AD and controls. RNA-seq, ATAC-seq, and pathway analyses were compared between ancestries in both AD and non-AD genes.
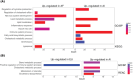
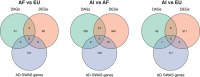
Results: Twelve AD genes were differentially expressed genes (DEGs) and/or accessible between ancestries, including ABI3, CTSB, and MS4A6A. A total of 5% of all genes had differential ancestral expression, but differences in accessibility were less than 1%. The DEGs were enriched in known AD pathways.
Discussion: This resource will be valuable in evaluating AD in admixed populations and other neurological disorders and understanding the AD risk differences between populations.
Highlights: First comparison of the genomics of AI, AF, and EU microglia. Report differences in expression and accessibility of AD genes between ancestries. Ancestral expression differences are greater than differences in accessibility. Good transcriptome correlation was seen between brain and iPSC-derived microglia. Differentially expressed AD genes were in known AD pathways.
Keywords: ATAC‐seq; Alzheimer's disease; RNA‐seq; diversity; genetic ancestry; genetic regulatory architecture; iPSC‐derived microglia.
© 2025 The Author(s). Alzheimer's & Dementia published by Wiley Periodicals LLC on behalf of Alzheimer's Association.
Conflict of interest statement
The authors declare nothing to disclose or competing interests. Author disclosures are available in the Supporting Information.
Figures

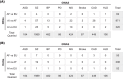
References
Publication types
MeSH terms
Supplementary concepts
Grants and funding
- U01-AG052410/AG/NIA NIH HHS/United States
- U19 AG074865/AG/NIA NIH HHS/United States
- Alzheimer Association, Zenith Award
- U01-AG066767/AG/NIA NIH HHS/United States
- RF1 AG059018/AG/NIA NIH HHS/United States
- R01 AG072547/AG/NIA NIH HHS/United States
- U01 AG052410/AG/NIA NIH HHS/United States
- U01 AG066767/AG/NIA NIH HHS/United States
- R01 AG070864/AG/NIA NIH HHS/United States
- RF1- AG059018/AG/NIA NIH HHS/United States
- U01-AG072579/AG/NIA NIH HHS/United States
- U01 AG072579/AG/NIA NIH HHS/United States
- R56-AG072547/AG/NIA NIH HHS/United States
- R01-AG070864/AG/NIA NIH HHS/United States
- R56 AG072547/AG/NIA NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

