Impairment of endocytosis-related factors FNBP1L, ARHGAP24, and ATP6V1B1 increases HIV-1 entry into dendritic cells
- PMID: 40029073
- PMCID: PMC11998494
- DOI: 10.1128/jvi.02066-24
Impairment of endocytosis-related factors FNBP1L, ARHGAP24, and ATP6V1B1 increases HIV-1 entry into dendritic cells
Abstract
HIV-1 infects several types of CD4+ cells. Among these, dendritic cells (DCs) are considered one of the first to encounter the virus upon sexual transmission. Expression of several restriction factors, of which SAMHD1 is well known, limits productive infection. Still, DCs are essential players in shaping adaptive immune responses that contribute heavily to the pathogenesis of HIV. Here, we set out to identify other factors that potentially contribute to the resistance of dendritic cells to HIV infection. Since endocytosis and the cytoskeleton impact HIV infection, we have put special emphasis on proteins implied in these pathways. In a selective, shRNA-mediated knockdown screen in primary monocyte-derived dendritic cells (MDDCs) infected with HIV in the presence of SAMHD1-disactivating Vpx containing virus-like particles, three proteins hampering HIV-1 infection were identified: FNBP1L, ARHGAP24, and ATP6V1B1. Findings of our research indicate that upon blocking of factors involved in endocytosis, increased viral entry is observed providing supportive evidence for endocytosis mostly being a dead-end entry pathway for HIV infection of MDDCs. Additional experiments show that changes in the cytoskeleton and endosomal pH that lead to impaired fluid-phase endocytosis and phagocytosis are responsible for these shifts in the phenotype observed.IMPORTANCEUnderstanding how HIV-1 interacts with dendritic cells (DCs) is pivotal in deciphering early viral transmission and immune evasion but is subject to a long-standing controversy in HIV virology. Therefore, the identification of endocytosis-related host factors as barriers to productive infection in DCs emphasizes the role of endocytosis as a restrictive pathway for viral entry. By disrupting these processes, we highlight a shift in the cellular environment that could influence viral entry and transmission. These findings challenge existing models of HIV-1 entry into DCs. New insights into how cellular pathways limit viral spread have implications for the development of strategies aimed to curb viral dissemination and reservoir formation. Whether the knockdown of the proteins described simply augments the efficiency of infection via existing pathways or opens additional routes for HIV-1 entry remains to be investigated.
Keywords: ARHGAP24; ATP6V1B1; FNBP1L; HIV-1; cytoskeleton; dendritic cells; endocytosis; viral entry.
Conflict of interest statement
The authors declare no conflict of interest.
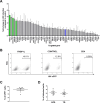
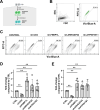
Figures





References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

