CTCF-mediated 3D chromatin sets up the gene expression program in the male germline
- PMID: 40033153
- PMCID: PMC12263338
- DOI: 10.1038/s41594-025-01482-z
CTCF-mediated 3D chromatin sets up the gene expression program in the male germline
Abstract
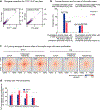
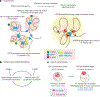
Spermatogenesis is a unidirectional differentiation process that generates haploid sperm, but how the gene expression program that directs this process is established is largely unknown. Here we determine the high-resolution three-dimensional (3D) chromatin architecture of mouse male germ cells during spermatogenesis and show that CTCF-mediated 3D chromatin dictates the gene expression program required for spermatogenesis. In undifferentiated spermatogonia, CTCF-mediated chromatin interactions between meiosis-specific super-enhancers (SEs) and their target genes precede activation of these SEs on autosomes. These meiotic SEs recruit the master transcription factor A-MYB (MYBL1) in meiotic spermatocytes, which strengthens their 3D contacts and instructs a burst of meiotic gene expression. We also find that at the mitosis-to-meiosis transition, the germline-specific Polycomb protein SCML2 facilitates the resolution of chromatin loops that are specific to mitotic spermatogonia. Moreover, SCML2 and A-MYB help shape the unique 3D chromatin organization of sex chromosomes during meiotic sex chromosome inactivation. We propose that CTCF-mediated 3D chromatin organization regulates epigenetic priming that directs unidirectional differentiation, thereby determining the cellular identity of the male germline.
© 2025. The Author(s), under exclusive licence to Springer Nature America, Inc.
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures














Update of
-
CTCF-mediated 3D chromatin predetermines the gene expression program in the male germline.bioRxiv [Preprint]. 2023 Dec 2:2023.11.30.569508. doi: 10.1101/2023.11.30.569508. bioRxiv. 2023. Update in: Nat Struct Mol Biol. 2025 Jul;32(7):1227-1240. doi: 10.1038/s41594-025-01482-z. PMID: 38076840 Free PMC article. Updated. Preprint.
References
-
- Zheng H & Xie W The role of 3D genome organization in development and cell differentiation. Nat. Rev. Mol. Cell Biol 20, 535–550 (2019). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

