Unveiling ac4C modification pattern: a prospective target for improving the response to immunotherapeutic strategies in melanoma
- PMID: 40050821
- PMCID: PMC11887236
- DOI: 10.1186/s12967-025-06297-6
Unveiling ac4C modification pattern: a prospective target for improving the response to immunotherapeutic strategies in melanoma
Abstract
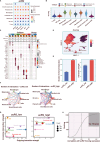
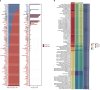
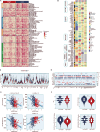
Emerging evidence has confirmed the inextricable connection between N4-acetylcytidine (ac4C) mRNA modification and the clinical characteristics of malignancies. Nonetheless, it is uncertain whether and how ac4C mRNA modification patterns affect clinical outcomes in melanoma patients. This research integrated single-cell sequencing data and transcriptomics to pinpoint ac4C-related genes (acRG) linked to melanoma progression and evaluate their clinical implications. Cells with elevated acRG score were predominantly located within the melanocytes cluster. Intercellular communications between melanocytes and other cell subtypes were markedly strengthened in the acRG-high group. We developed and confirmed an excellent acRG-related signature (acRGS) utilizing a comprehensive set of 101 algorithm combinations derived from 10 machine learning algorithms. Hereby, the acRGS, including MYO10, ZNF667, MRAS, SCO2, MAPK10, PNMA6A, KPNA2, NT5DC2, BAIAP2L2 and NDST3, delineated ac4C-associated mRNA modification patterns in melanoma. The acRGS possesses distinctly superior performance to 120 previously reported signatures in melanoma and could predict the overall survival of melanoma patients across four external datasets. The substantial associations among immune checkpoint genes, immune cell infiltration, and tumor mutation burden with acRGS indicate that acRGS is helpful in identifying melanoma patients who are sensitive to immunotherapy. Besides, we confirmed that MYO10 was mainly overexpressed in melanoma tissues, and elevated MYO10 was positively correlated with malignant phenotypes and unfavorable prognosis in melanoma patients. Silencing MYO10 expression inhibited melanoma cell proliferation, migration and invasion in vitro as well as tumor growth in vivo. Taken together, the acRGS could function as a reliable and prospective tool to improve the clinical prognosis for melanoma individuals.
Keywords: Immunotherapy; MYO10; Machine learning; Melanoma; Tumor microenvironment; ac4C modification.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Ethics Committee of the First Affiliated Hospital of Nanjing Medical University (30 August 2022/ No.2022-SR-465). Informed consent has been obtained from all participants involved in the study. All animal procedures adhered to the ethical guidelines of the Animal Experiments Committee at Nanjing Medical University (IACUC-2403031). Consent for publication: Consent to publish was obtained from the study participants. Competing interests: The authors declare that they have no competing interests.
Figures







References
-
- Long GV, Swetter SM, Menzies AM, Gershenwald JE, Scolyer RA. Cutaneous melanoma. Lancet. 2023;402(10400):485–502. - PubMed
-
- Ascierto PA, Dréno B, Larkin J, Ribas A, Liszkay G, Maio M, Mandalà M, Demidov L, Stroyakovskiy D, Thomas L, et al. 5-Year outcomes with Cobimetinib plus Vemurafenib in BRAFV600 Mutation-Positive advanced melanoma: extended Follow-up of the cobrim study. Clin Cancer Res. 2021;27(19):5225–35. - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

