Profiling the response of individual gut microbes to free fatty acids (FFAs) found in human milk
- PMID: 40051690
- PMCID: PMC11884519
- DOI: 10.1016/j.jff.2025.106664
Profiling the response of individual gut microbes to free fatty acids (FFAs) found in human milk
Abstract
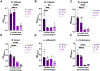
Preterm infants have an immature intestinal environment featuring microbial dysbiosis. Human milk can improve the composition of the neonatal gut microbiome by supporting commensal species. Milk free fatty acids (FFAs) provide nutritional energy, participate in endogenous signaling, and exert antimicrobial effects. This study examined the growth of individual commensal and pathobiont microbes in response to unesterified unsaturated FFAs found in milk: oleic, linoleic, arachidonic, and docosahexaenoic acid. Select species of commensal and pathobiont genera (Bifidobacterium, Lactobacillus, Streptococcus, Staphylococcus, Enterococcus, Acinetobacter, Pseudomonas, Escherichia, and Klebsiella) were cultured with FFAs. The growth of all commensals, except for L. johnsonii, was significantly inhibited by the highest concentration (1 %) of all FFAs. L. johnsonii was only inhibited by arachidonic acid. In contrast, suppression of pathobionts in response to FFAs was less pronounced. Higher concentrations (0.1 %, 1 %) of docosahexaenoic acid significantly inhibited the growth of five of eight pathobionts. Meanwhile, for oleic, linoleic, and arachidonic acid, only two of eight pathobionts were significantly affected. Intriguingly, the effects for these FFAs were highly complex. For example, S. agalactiae growth was enhanced with 1 % oleic acid but suppressed at 0.01 %; however, the effects were directionally opposite for linoleic acid, i.e., suppressed at 1 % but enhanced at 0.01 %. Our genome analyses suggest that pathobiont survival may be related to the number of gene copies for fatty acid transporters. Overall, the effect of FFAs was dose-dependent and species-specific, where commensal growth was broadly inhibited while pathobionts were either unaffected or exhibited complex, bi-directional responses.
Keywords: Bifidobacterium; Diet; Enterobacter; Enterococcus; Formula; Free fatty acid; Human breast milk; Intestine; Klebsiella; Lactobacillus; Microbiome; Pathogens; Premature infant.
Conflict of interest statement
Disclosures The authors declare that the research was conducted without any commercial or financial relationships that could be construed as a potential conflict of interest. Declaration of competing interest The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures







References
-
- Adlerberth I, & Wold A (2009). Establishment of the gut microbiota in Western infants. Acta Paediatrica, 98(2), 229–238. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
