This is a preprint.
Class I histone deacetylases catalyze lysine lactylation
- PMID: 40060688
- PMCID: PMC11888385
- DOI: 10.1101/2025.02.25.640220
Class I histone deacetylases catalyze lysine lactylation
Update in
-
Class I histone deacetylases catalyze lysine lactylation.J Biol Chem. 2025 Oct;301(10):110602. doi: 10.1016/j.jbc.2025.110602. Epub 2025 Aug 18. J Biol Chem. 2025. PMID: 40835008 Free PMC article.
Abstract
Metabolism and post-translational modifications (PTMs) are intrinsically linked and the number of identified metabolites that can covalently modify proteins continues to increase. This metabolism/PTM crosstalk is especially true for lactate, the product of anaerobic metabolism following glycolysis. Lactate forms an amide bond with the ε-amino group of lysine, a modification known as lysine lactylation, or Kla. Multiple independent mechanisms have been proposed in the formation of Kla, including p300/CBP-dependent transfer from lactyl-CoA, via a high-energy intermediate lactoylglutathione species that non-enzymatically lactylates proteins, and several enzymes are reported to have lactyl transferase capability. We recently discovered that class I histone deacetylases (HDACs) 1, 2, and 3 can all reverse their canonical chemical reaction to catalyze lysine β-hydroxybutyrylation. Here we tested the hypothesis that HDACs can also catalyze Kla formation. Using biochemical, pharmacological, and genetic approaches, we found that HDAC-catalyzed lysine lactylation accounts for the majority of Kla formation in cells. Dialysis experiments confirm this is a reversible reaction that depends on lactate concentration. We also directly quantified intracellular lactyl-CoA and found that Kla abundance can be uncoupled from lactyl-CoA levels. Therefore, we propose a model in which the majority of Kla is formed through enzymatic addition of lactate by HDACs 1, 2, and 3.
Conflict of interest statement
Declaration of Interests The authors declare no competing interests.
Figures

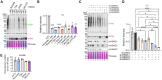


References
-
- Verdin E., and Ott M. (2015) 50 years of protein acetylation: from gene regulation to epigenetics, metabolism and beyond Nat Rev Mol Cell Biol 16, 258–264 - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
