Spatial 3D genome organization reveals intratumor heterogeneity in primary glioblastoma samples
- PMID: 40073147
- PMCID: PMC11900876
- DOI: 10.1126/sciadv.adn2830
Spatial 3D genome organization reveals intratumor heterogeneity in primary glioblastoma samples
Abstract
Glioblastoma (GBM) is the most prevalent malignant brain tumor with poor prognosis. Although chromatin intratumoral heterogeneity is a characteristic feature of GBM, most current studies are conducted at a single tumor site. To investigate the GBM-specific 3D genome organization and its heterogeneity, we conducted Hi-C experiments in 21 GBM samples from nine patients, along with three normal brain samples. We identified genome subcompartmentalization and chromatin interactions specific to GBM, as well as extensive intertumoral and intratumoral heterogeneity at these levels. We identified copy number variants (CNVs) and structural variations (SVs) and demonstrated how they disrupted 3D genome structures. SVs could not only induce enhancer hijacking but also cause the loss of enhancers to the same gene, both of which contributed to gene dysregulation. Our findings provide insights into the GBM-specific 3D genome organization and the intratumoral heterogeneity of this organization and open avenues for understanding this devastating disease.
Figures
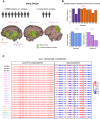

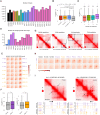


References
-
- Tan A. C., Ashley D. M., Lopez G. Y., Malinzak M., Friedman H. S., Khasraw M., Management of glioblastoma: State of the art and future directions. CA Cancer J. Clin. 70, 299–312 (2020). - PubMed

