KinasePred: A Computational Tool for Small-Molecule Kinase Target Prediction
- PMID: 40076779
- PMCID: PMC11900317
- DOI: 10.3390/ijms26052157
KinasePred: A Computational Tool for Small-Molecule Kinase Target Prediction
Abstract
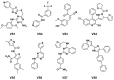
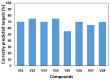
Protein kinases are key regulators of cellular processes and critical therapeutic targets in diseases like cancer, making them a focal point for drug discovery efforts. In this context, we developed KinasePred, a robust computational workflow that combines machine learning and explainable artificial intelligence to predict the kinase activity of small molecules while providing detailed insights into the structural features driving ligand-target interactions. Our kinase-family predictive tool demonstrated significant performance, validated through virtual screening, where it successfully identified six kinase inhibitors. Target-focused operational models were subsequently developed to refine target-specific predictions, enabling the identification of molecular determinants of kinase selectivity. This integrated framework not only accelerates the screening and identification of kinase-targeting compounds but also supports broader applications in target identification, polypharmacology studies, and off-target effect analysis, providing a versatile tool for streamlining the drug discovery process.
Keywords: kinase; machine learning; virtual screening.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
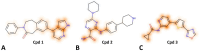
Similar articles
-
The Development and Application of KinomePro-DL: A Deep Learning Based Online Small Molecule Kinome Selectivity Profiling Prediction Platform.J Chem Inf Model. 2024 Oct 14;64(19):7273-7290. doi: 10.1021/acs.jcim.4c00595. Epub 2024 Sep 25. J Chem Inf Model. 2024. PMID: 39320984
-
Data structures for computational compound promiscuity analysis and exemplary applications to inhibitors of the human kinome.J Comput Aided Mol Des. 2020 Jan;34(1):1-10. doi: 10.1007/s10822-019-00266-0. Epub 2019 Dec 2. J Comput Aided Mol Des. 2020. PMID: 31792884
-
Application of Free-Wilson selectivity analysis for combinatorial library design.Methods Mol Biol. 2011;685:91-109. doi: 10.1007/978-1-60761-931-4_5. Methods Mol Biol. 2011. PMID: 20981520
-
Quantitative proteomics of kinase inhibitor targets and mechanisms.ACS Chem Biol. 2015 Jan 16;10(1):201-12. doi: 10.1021/cb5008794. Epub 2014 Dec 17. ACS Chem Biol. 2015. PMID: 25474541 Review.
-
Leveraging Structural Diversity and Allosteric Regulatory Mechanisms of Protein Kinases in the Discovery of Small Molecule Inhibitors.Curr Med Chem. 2017;24(42):4838-4872. doi: 10.2174/0929867323666161006113418. Curr Med Chem. 2017. PMID: 27719654 Review.
References
-
- Song J., Wang H., Wang J., Leier A., Marquez-Lago T., Yang B., Zhang Z., Akutsu T., Webb G.I., Daly R.J. PhosphoPredict: A Bioinformatics Tool for Prediction of Human Kinase-Specific Phosphorylation Substrates and Sites by Integrating Heterogeneous Feature Selection. Sci. Rep. 2017;7:6862. doi: 10.1038/s41598-017-07199-4. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

